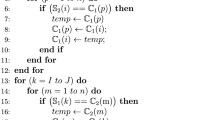Abstract
In the Steiner Tree Problem in Graphs, a subset of nodes, called terminals, must be efficiently interconnected. The graph is undirected and weighted, and candidate solutions can include additional nodes called Steiner vertices. The problem is NP-complete, and optimization algorithms based on population metaheuristics, e.g., Genetic Algorithms (GAs), have been proposed. However, traditional recombination operators may produce inefficient solutions for graph-based problems. We propose a new recombination operator for the Steiner Tree Problem based on the graph representation. The new operator is a kind of partition crossover: it breaks the graph formed by the union of two parents solutions and decomposes the evaluation function by finding connected subgraphs. We also investigate two soft-repair operators that produce small changes in candidate solutions: an MST transformation and a pruning repair. The GA with the proposed crossover operator was able to find the global optimum solution for all tested instances and has a success rate of 28% in the worst case. The experiments with the proposed crossover presented a quick convergence to optimal solutions, and an average solution cost lower in most cases than other approaches.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Steiner Tree Problems (STP) are relevant in the combinatorial optimization field [9]. In STP, a minimum special tree should be found in a weighted graph G(V, E), where an edge \(e_{ij} \in E\) represents the relationship between vertices \(v_i\) and \(v_j \in V\), and a function \(c: E \rightarrow \mathbb {R}^{+}_{*}\) maps each edge to a positive value. The graph is also undirected, i.e., we can reach a node \(v_j\) from \(v_i\) and vice-versa by the same edge and cost. Moreover, suppose there is a subset of requested vertices \(Z \subset V\), also called terminals.
The Steiner Tree Problem in Graphs (STPG) seeks the Steiner Minimum Tree (SMT), a subgraph of G that connects all the terminals nodes in Z and minimize Eq. 1, i.e., has the minimum cost.
A Steiner tree (ST) is a candidate solution for the problem with \(Z \subset V'\) and \(E' \subset E\) forming a tree \(T(V', E')\). Some non-terminal vertices from the set \(V - Z\) might appear in \(V'\) to assure connectivity - they are Steiner vertices. An ST must follow these constraints: 1) The terminal nodes must be connected; 2) Any cycle must be broken to reduce the solution cost; 3) Leaf nodes that are not terminals must be removed, reducing the solution cost without lost connectivity.
Hamiki (1971) [4] first described the SPTG for a finite graph with vertices and well-defined edges. The STPG is a well-known NP-complete problem [6] in the combinatorial optimization field. It means no known algorithm assures that deterministically can find the best possible solution in a polynomial time for the general case. As a consequence, heuristics and metaheuristics have been proposed for the STPG.
The STPG can model a comprehensive range of practical problems. Several studies describe applications in the telecommunication and related fields as Multicast Routing Problems and Wireless Sensor Networks. However, it is also applicable in supply chain management, VSLI circuit projects, genetic phylogeny, and drug repositioning. Due to its applicability, several studies attempted to find better solutions for the STPG.
Ljubic (2021) [9] describes different approaches for special cases of the STPG: Lagrangian relaxations, dual-ascent methods, heuristics, approximate and exact algorithms. The author in [9] pointed out that metaheuristics also has been developed, like genetic algorithms (GA), ant colony optimization, and particle swarm optimization. However, metaheuristics are not competitive in many practical Steiner Tree benchmark problems where traditional approaches have been applied. The worst results, when compared to conventional approaches, can be partially credited to the use of transformation operators that end up producing inefficient or unfeasible solutions in graph-based representations of the problem.
In addition, the interaction between decision variables is ignored in black-box optimization. Alternatively, in gray-box optimization, the structure of the problems is efficiently explored by transformation operators to produce better solutions [14]. Partition Crossover (PX) operators explore the interaction between decision variables in problems with graph-based representation to recombine solutions efficiently. PX operators break the union graph formed by two parents’ solutions and decompose the problem’s evaluation function.
For instance, the Generalized Partition Crossover (GPX) improved the solution cost for the Traveling Salesman Problem (TSP) when it was combined with the state-of-art Lin-Kernighan-Helsgaum heuristic [13]. This operator is respectful because the offspring inherits all common edges from its parents. It also transmits alleles, which means all edges present in the offspring can be found in the parents. Moreover, when two local optima are recombined, the offspring is also local optima.
Accordingly to Raidl et al. (2003) [11], an efficient crossover operator for tree-graph problems is expected to be respectful and transmit alleles. In this paper, we propose a new recombination operator for the STPG based on partition crossover. It is important to observe that PX operators were not previously proposed for Steiner Tree Problems. In addition, we use two soft-repair operations to repair unfeasible solutions eventually generated by recombination. The proposed operator is compared to a Prim’s random-based crossover. The GA with the proposed recombination and repair operators is also compared to a classical GA using binary chromosome representation.
This paper is organized as follows: Sect. 2 describes GA approaches for the STPG and some expected features for an efficient crossover operator. Section 3 explains our partition crossover for Steiner Trees. Section 4 describes the experiments, Subsect. 4.1 gives more detail about GA tested, and Subsect. 4.2 discusses results. Finally, a conclusion is given in Sect. 5.
2 Related Works
In two early studies, Kapsalis et al. (1993) [5] and Esbensen (1995) [3] proposed a GA with a binary chromosome to encode an ST. The binary chromosome indicates if a non-terminal node (from the set \(V - Z\)) belongs to an ST. If it belongs, it is set the value one to the corresponding node. Otherwise, it will set a value of zero if the node does not belong. The algorithms in [5] and [3] differ by the process of decoding the ST from the chromosome.
The decoding procedure in [5] first identifies the Steiner vertices from the chromosome and the terminals nodes that form the set of nodes \(V' \subset V\) for the ST. Then, it adds an edge \( e_{ij} = (v_i, v_j) \in E\) to the subgraph \(G'\) if both incidents vertices \(v_i\) and \(v_j\) belongs to \(V'\). If so, the edge will also belong to \(G'\). Finally, the ST is the Minimum Spanning Tree (MST) from the subgraph \(G'\).
The decoding procedure adopted by [5] can lead to disconnected individuals. However, their fitness evaluation receives a penalization proportionally to the number of disconnected components. They also used traditional GA operators such as 1 point crossover and bit-flip mutation. The GA found the best solution for all OR-Library B class instances in at least one algorithm run. However, the authors applied a diverse range of parameters configuration to achieve the results, and the GA required a long running time.
Esbensen (1995) [3] embodied the Distance Network Heuristic (DNH) [7] to avert disconnected solutions in his decoding procedure. The heuristic uses the distance graph (representing all paths and distances between every pair of vertices) obtained by the Floyd-Wharsall algorithm [2]. Even though it is computed once, its time complexity of \(O(n^3)\) might prevent its use for large graphs. The decoding procedure computes the ST by applying the heuristic over the subgraph \(G'\) induced by the chromosome subset of vertices \(V'\).
The author in [3] applied his GA in C, D, and E instance classes from OR-Library [1]. The author also used reduction techniques to reduce the size of the instance graphs. Reduction techniques try to identify edges and vertices that always will belong to an ST—contracting those pairs of vertices and edges—or identifying those that will never be part of an ST and remove them. However, the GA achieved a near-optimum solution and took a long time running.
Raidl et al. (2003) [11] presented an Edge Set encoding for tree-based optimization problems. In this approach, just the set of edges represents the tree. They tested it considering different tree-based problems, but the adaptation to STPG is simple. They also proposed three randomized strategies for population initialization, based on Prim [10], Kruskal [8], and random walk algorithms.
The random walk starts at an arbitrary node \(v_{start}\) and moves a particle to some adjacent node \(v_k\). The edge \((v_{start}, v_k)\) is selected to compose the resulting tree. Then the particle moves to another adjacent node of \(v_k\) and inserts a new edge into the resulting tree if the selected edge does not insert a cycle in the tree. This procedure is repeated until all terminal nodes belong to the individual. The authors in [11] proved that this procedure does not favor a specific tree topology.
Prim (1957) [10] and Kruskal (1956) [8] have proposed two well-known algorithms to compute the MST from a graph. These algorithms greedily choose the lowest cost edges to form the MST, but they differ in selecting such edges. Kruskal’s algorithm looks at the entire edges’ set and selects one even though it does not connect with the previously chosen. Therefore, this algorithm uses a disjoint set data structure to ensure that the selected edges will not form a cycle. Instead, Prim’s algorithm starts from a node \(v_{0}\), considers its incident edges as eligible, and chooses some edge \((v_0, v_j)\) with the lowest cost. The new incident edges from \(v_j\) will be eligible too. The set of eligible edges keeps growing until all graph edges are reached.
As a procedure to population initialization, the authors in [11] proposed a variation of Prim and Kruskal algorithms to select a new edge randomly instead of in a greedy way. Furthermore, the authors claimed that these procedures favor a star topology in the individuals. Hence, they named these variations as PrimRST and KruskalRST.
Also, in [11] the initialization procedures are adapted as recombination procedures by simply applying them to the union graph of the two candidate solutions. In addition, they stated that other recombination strategies are possible depending on what to do with the shared edges, transmitting them directly to the offspring or not. However, they conclude that recombination operators for spanning trees should always include the common edges in the offspring.
The authors in [11] stated that these operators offer locality (small changes in the genotype imply small changes in the phenotype) and heritability (the offspring consist mainly of alleles of its parents). These are two of many properties expected for proper encoding. However, to the best of our knowledge, those recombination strategies were not evaluated for the STPG case. Additionally, there is no formal demonstration that PrimRST always includes all the common edges between two individuals chosen for recombination.
In this paper, we propose a crossover operator for ST based on PX. It generates two offspring, which will have: the shared edges from both parents, the select edges from recombinant partitions, and the remaining edges from non-recombinant partitions, respectively, from each parent.
The principles behind our operator come from the PX for the Traveling Salesman Problem (TSP) proposed in [12, 13, 15], and [16]. The original Generalized Partition Crossover (GPX) was designed to recombine two Hamiltonian cycles. Under these circumstances, a new recombination operator is necessary for the Steiner Tree, i.e., recombine tree graph solutions.
The GPX takes two individuals (Hamiltonian cycles) for recombination. Then, it joins the edges from each individual in a union graph \(G_u\). The vertices with a degree equal to 2 are removed, and their incident edges are transferred directly to the offspring. The remaining edges form candidate partitions for recombination. One can think each partition is the union of two paths, each one from a parent.
The recombinant partition must ensure that it can ensemble a cycle in the offspring if either inner path is chosen. It is made by checking the portal and simplified graph for each parent graph. A simplified graph reduces a path to a single edge. A portal vertex connects a partition to another through common edges. So it will be easy to verify if the path from both parents connects to the same portal vertex. Nevertheless, vertices with a degree equal to 4 are split by inserting a ghost vertex. The edges are then divided between each vertex–the original and the ghost one.
An example of PXST crossover between two ST, \(T_a\) and \(T_b\), showed in Figs. 1a and 1b. Figure 1c shows the resulting graph after removing the common edges from the union \(T_a \cup T_b\); the resulting graph has only one connected component. Figure 1d shows the common elements (vertices and edges) between \(T_a\) and \(T_b\). They will be used to explore more partitions. PXST represents common elements as a disjoint set, where vertices are linked if they share a common edge (steps 1, 2, and 3, of Algorithm 1). Figure 1e shows two cycles identified in the first iteration. The distinct paths of non-common edges are connected to the same components of common elements. Then, we can choose the best path. The selected path is added to the common elements, and two more cycles are then identified in the second iteration (Fig. 1f). Again, the respective paths for each parent are compared, and the best paths are chosen. Figure 1g shows the offspring. All the partitions are feasible for recombination in this example, but this may not be the case for some pairs of solutions (ST).
3 Proposed Operator
As mentioned before, the original GPX is not suitable for the STPG case due to differences in candidate solutions’ topology. Algorithm 1 shows the proposed recombination operator based on partition crossover (PXST).
Both operators (the original and the proposed one) keep some resemblance. The offspring will inherit all the common edges between the parents. Uncommon edges will form partitions, and some tests will identify the comparable ones, also called recombinant partitions. For such partitions, those with the lowest cost will compose the offspring. Moreover, if there are k recombining components, the operator will generate the best of \(2^k\) potential offspring.
Furthermore, the PXST’s operation can be divided into four phases: identifying common elements (vertices and edges); identifying partitions candidates to recombination; matching, selecting, and updating the recombinant partitions; handle partitions unfeasible to recombination and return the offspring. Each phase will be detailed as follows.
a) Identifying Common Elements. The common vertices and edges between two ST, play an essential role in the PXST. They work as a cutting point, increasing the number of partitions and will help to identify recombinant partitions.
The vertices from the union graph between two Hamiltonian cycles will have degrees equal to 2, 3, or 4, and GPX explores it. First, the operator removes the vertices with a degree equal to 2. The vertices with a degree equal to 3 belong to some partition. Moreover, the insertion of a ghost vertex will separate the edges adjacent to vertices with a degree equal to 4, leading to more partitions.
Unlike, the ST inner vertices do not have such degree constraints. An inner node can have any number of adjacent edges, and only the leaves must have a degree equal to one. Consequently, the union graph between two ST solutions might have more connections within its vertices, leading to fewer independent components.
Figures 1a and 1b show two examples of ST, \(T_a\) and \(T_b\). Figure 1c shows why the strategy adopted by GPX is not suitable for the STPG. Figure 1c the resulting graph obtained by removing the common edges from the union between \(T_a\) and \(T_b\) has only one connected component. In this simple case, choosing either one or another parent’s edges did not result in a new individual.
Figure 1d shows the common edges and vertices between \(T_a\) and \(T_b\). Notice that the vertices 15, 61, and 47 will act like cutting points, breaking the component shown in Fig. 1c in smaller partitions.

Moreover, there are two sets of common edges: one formed by the edge \(\{(5, 9)\}\) and the other formed by the edges \(\{(55, 12), (12, 11), (23, 11), (17, 11)\}\). Also, notice how candidate partitions connect to these sets by different vertices. One first test to identify a recombinant partition might be to check when two distinct partitions (one from each parent) connect to the same components of common elements, i.e., a set of connected edges or an isolated vertex.
To keep track of such common elements, PXST uses a disjoint set (DS) data structure. It allows three operations: the “make a set” initializes the disjoint set; the “find” operation returns the same representative element for any other element in the set; the “union” operation joins two sets and defines a new element to represent the new set. An efficient implementation of the union operation (using union by rank and path compression) has \(O(m \log n)\) time complexity [2].
Step 1 from Algorithm 1 firstly identifies the common vertices between the parent’s solutions. Then, step 2 initializes DS by creating a unitary set for all common vertices. Finally, DS’s initial unitary sets are jointed when a common edge connects two distinct vertices (step 3). Notice that the set of common edges (showed in Fig. 1d) will be represented as a disjoint set, i.e., unique key-vertex.
Another detail is that some individuals might have leaf nodes that are not terminals. Some of these leaf nodes might belong to just one of the parent’s solutions. Moreover, such leaf nodes will become a portal vertex for one partition. Since checking portal vertices will allow identifying recombinant partition, such leaves will be included in the DS, although they may not be common (step 1).
b) Identifying Candidate Partitions to Recombination. The PXST identifies partitions directly on the graphs \(T_a^*\) e \(T_b^*\) (step 4). They are the resulting graphs from deleting the common edges from \(T_a\) and \(T_b\), respectively. Removing an edge from a tree results in two subtrees. Therefore, \(T_a^*\) and \(T_b^*\) are compound by many subtrees. However, those subtrees are not partitions yet. They still will be divided into smaller parts taking the common vertices as cutting points.
A depth-first search looks for partitions in \(T_a^*\) e \(T_b^*\), separately (step 5). A partition is formed mainly by non-common elements, where all its internal vertices and edges belong to one of the solutions. Since the common vertices work as cutting points, where the partitions are breaking, these vertices are called portals. Figure 1 shows partitions only with two portal vertices. However, a partition with more than two portal vertices is possible when an internal node has a degree greater than two, i.e., the partition has a subtree topology.
c) Matching, Selecting, and Updating the Recombinant Partitions. For instance, consider two STs with common vertices \(u, v \in T_a \cap T_b\). Pretend that \(T_a\) has a partition with a path-like topology \(P_a\) and \(T_b\) has a \(P_b\) partition. Both \(P_a\) and \(P_b\) connect u and v by different edges. The union between \(P_a\) and \(P_b\) will form a cycle closing in the vertices u and v. Moreover, u and v are the portal vertices for these partitions. Alternatively, the portal vertices for the partitions are coincident, and one can select the partition with the lower cost.
However, Figs. 1e and 1f show that partitions can connect to the same component of common edges by different vertices. Then, the partition will not have coincident portal vertices. So, a more refined way to find recombinant partitions is needed.
One can argue that if we shrink all common edges to a single vertex, we might get the well-formed cycle as mentioned earlier. For example, in Fig. 1e, vertices 12 and 23 will become coincident, and the partitions will form a cycle between nodes 15 and 12 (or 23). In the same way, shrinking the edge (5, 9) will close another cycle with the path-like partitions up to node 17. Then, we will be able to compare such distinct paths.
Instead of shrinking common edges and checking the exact correspondence between portal vertices from distinct partitions, we check if they are mapped to the same representative nodes by the disjoint set (step 6). That indicates the partitions are connected to the same components of common elements (Fig. 1d), and they are interchangeable. Then, the operator can choose between the partitions without losing the offspring connectivity or adding a loop into it.
When a partition is liable for recombination, the disjoint set from each portal vertex can be joined, expanding the original disjoint set (common elements set). This update is helpful to discover new recombination opportunities, and it will be performed until there are no new recombinant components (step 9).
For instance, Fig. 1f shows other opportunities for recombination. In the first one, we can choose the path from node 61 up to 15 or from 61 to 55. However, to compare these paths, it is necessary to resolve the connectivity to node 15 first (shown in Fig. 1e). When we resolve the best connectivity up to node 15, this node will belong to the same disjoint set with nodes 12, 23, and 55. The portal vertices 15 and 55 will be mapped to the same arbitrary representative element, and the operator can decide the best path to node 61. A similar situation occurs to the partitions connecting node 47. Firstly, we must decide the best connectivity up to node 9 to include this node to the same disjoint set as portal vertex 12.
Some partitions might have equal cost even though they do not have the same edges. In such cases, the selection occurs randomly, with equal probability for each partition. When their total cost is different, PXST chooses the one with a lower cost. Thus, the partition’s selected edges will belong to both offspring.
d) Handle Unfeasible Partitions to Recombination and Return Offspring. By the end of this process, some non-recombinant partitions might persist for both parents. One alternative is to choose the remaining partitions from one of the parents with the lowest sum cost. In preliminary tests, this strategy leads to an early population stagnation and higher solution cost outcomes. Another option is to return two offspring, one for each parent’s remaining partitions (step 10). Therefore, we chose this last alternative as it showed to lead to better outcomes.
New ways for identifying recombinant partitions can further be developed. It may improve the operator’s efficiency and lead to better solutions.
4 Experiments
This section describes the experiments comparing the proposed operator (PXST), the PrimRST crossover operator [11] and a binary GA based on [5]. The section also describes soft-repair operators used in our algorithm. In the experiments, three algorithms were compared:
-
PXST+Repair(pruner, mst): GA with the proposed operator PXST (Algorithm 1), mutation that replaces edges randomly, and the pruning and MST repair strategies.
-
PrimRST+Repair(pruner, mst): GA with the PrimRST crossover operator [11] and pruning and MST repair strategies.
-
Binary GA: GA with binary chromosome [5]. It used a traditional 2 point crossover operator and bit-flip mutation.
4.1 Experimental Design
The algorithms were tested over 18 instances of class B from the OR-Library [1]. Table 1 presents the properties for each instance and statistics about the solution cost discussed later. In all three experiments, the same stop condition was adopted. The execution stops if the algorithm finds the global optimum solution, if the best solution is not improved within an interval of 500 fitness evaluations, or if the number of iterations reaches 4.000. However, this last condition was never reached.
The experiments with PXST and PrimRST differ only by the crossover operator. They have the same parameters and underlying mechanisms for selection (roulette) and mutation. An adjacency list data structure represents the ST. The population size is 100 individuals, initialized by a heuristic based on the random walk procedure presented in [11]. This initialization strategy generates only connected individuals.
The mutation by replacing a random edge (RRE) is presented in [11]. The operator removes a random edge from the tree producing two disconnected subtrees. Then, it reconnects them by inserting a new edge selected randomly from the problem instance graph. The mutation rate is fixed to 0.3. This operator produces a slight change in the ST since it replaces just one edge when possible.
The PrimRST crossover takes two individuals and joins them in a graph \(G_u\). Like the original Prim’s algorithm, the operator computes a spanning tree from \(G_u\) (see Sect. 2). Instead of taking the lowest cost edge, the operators insert a new edge randomly – of course, if the new edge does not form a cycle with the previously selected edges. The algorithm stops when all terminal vertices are inserted in the offspring or when all edges from \(G_u\) were processed.
The offspring’s edges generated by PrimRST came only from \(G_u\), i.e., the operator does not insert a new edge that does not belong to one of the parents. Some common edges will appear in the offspring, although there is no proof that all common edges will be transmitted every time. In that regard, both PrimRST and PXST transmit alleles (edges) from the parent to the offspring. However, PXST transmits all of the common edges to the offspring, and for PrimRST, this is not guaranteed. Moreover, PrimRST favors the edge exchange between the parents, while PXST looks for connected components.
PrimRST and PXST generate connected individuals. Still, some offspring might not be completely ST because some of them might have leaf nodes that are not terminal. Thereby, a pruning repair iteratively seeks and removes leaf nodes that are not terminal. It reduces the solution cost without losing the offspring’s connectivity. Thus, the pruning repair was constantly applied for all individuals over the generations (fitness evaluations).
In [5] and [3] a solution is obtained computing the MST of the induced subgraph from Steiner vertices represented in the chromosome. Similarly, an ST can be improved by replacing some of its edges with lowest cost edges that span the solution vertices, computed from the induced MST subgraph. For that reason, an MST-based repair is applied.
The MST-based repair operator takes one individual \(T_k(V_k, E_k)\) and determines a subgraph induced from its vertices \(V_k\). The induced subgraph has all edges from the individual plus the edges from the problem instance graph when both incident vertices are in the individual’s vertex set. Then, it applies Prim’s algorithm to compute the MST from the induced subgraph. This repair operator is used with a fixed rate of 0.3.
Since PXST and PrimRST operators do not produce disconnected individuals, repair operations to reconnect the candidate solution or a penalty function (as in [5]) are useless.
The binary GA is based on [5] (see Sect. 2). A binary chromosome represents the Steiner vertices that belong to some ST. A decoding procedure based on computing the MST tree from the induced graph does not assure the candidate solution’s connectivity. Therefore, a penalty function increases the fitness of infeasible solutions (disconnected ones). The GA used a population size of 100 individuals. The binary chromosome is randomly initialized, and the genetic operators are roulette selection, a traditional crossover operator with 2 points cut, and a mutation by bit flip by a rate of 0,2.
The operators and GA implementations were in Python language, version 3.7.9, and the code is available publicly.Footnote 1 Experiments were conducted in a virtual machine set up in Google CloudTM platform, using Linux Ubuntu OS (version 18.4), 32 GB of RAM, and CPU with 8 cores Intel Cascade Lake. Each experiment runs 50 times, with a seed randomly defined by the system.
4.2 Experimental Results and Analysis
Table 1 presents the instances graphs features alongside the average and standard deviation for the best solution cost found in all executions. The values in bold indicate the lower average cost. Table 2 shows the average and standard deviation of execution time (in seconds) and the success rate (SR), i.e., the proportion of runs where the algorithm found the global optimum solution.
For the average cost and execution time results, we applied the Wilcoxon signed-rank test to verify if the differences in the samples are statistically significant. We adopted a significance level of \(\alpha =0.05\).
The GA with PXST found the global optimum solution for all instances in at least one run. This GA found the global optimum solution for all execution in five instances (B1, B2, B3, B8, and B9). For the other six cases (B11, B13, B14, B15, B16, and B18), the GA obtained the lowest average solution cost, as showed in Table 1. Moreover, GA with PXST has an SR in the worst case of 28% (instance B15), as shown in Table 2. In all other cases, it has an SR above 30%, indicating certain robustness of the algorithm.
The GA with PrimRST found the global optimum solution in all runs for seven instances: B2, B3, B5, B7, B9, B10, and B17. It obtained the lowest average solution cost for the B4 instance. Significant improvements are observed for B4, B10, and B17. This GA could not find the optimum solution in any run for B14, B15, and B16 and obtained a low SR for instances B6 (24%) and B12 (2%) (Table 2).
The statistical test does not indicate significant differences for the best cost sample to B1, B2, B3, B5, B7, B8, B9, B11, and B18 between PXST and PrimRST experiments. For example, GA with PrimRST found the global optimum solution, for instance, B5 in all runs (SR 100%) – note that the average best cost is equal to the global optimum, and the standard variation is zero. In contrast, GA with PXST obtained 94% of the SR and an average best cost of 61.10 and a standard deviation of 0.46. However, the test indicates that the difference observed in the best cost sample was not significant for the B5 instance.
Binary GA obtained a lower best cost average for instances B6 and B12. However, the differences for best cost are statistically significant just for B6. All of the three GA solved the instance B3 in all runs. For all other 15 instances, the best cost average for binary GA was worse than the GA with PXST. This result can be credited to the use of specific operators for tree-graph problems rather than traditional operators that produce infeasible (disconnected) solutions.
The GA with PXST was faster for fourteen instances (see Table 2). Moreover, GA with PXST was, in general, faster than PrimRST for those instances where the average best cost was similar, i.e., differences were not statistically significant. That is the case for B1, B2, B3, B5, B8, B9, and B11 (except B7). This result is because the crossover PXST is greedy and converges quickly to a local optimum solution.
The GA with PrimRST was faster when it found the global optimum solution in all runs (B7, B10, and B17) and in B4, where the SR is 86%. PXST and PrimRST GA took less than a second on average to solve some instances (see B2, B3, B7, B8, B9, and B10).
As expected, binary GA usually took a long time to run on average, as showed in Table 2. For B6, binary GA was faster than the other two GAs. Compared with GA with PXST, the execution time is similar (not statistically significant) for B6, B15 and B18, and worse for all other instances. Both PXST and PrimRST operators do not rely on high complexity cost heuristic as [3]. Hence, they might be suitable for large instance graphs in an appropriate execution time.
5 Conclusions
In this paper, we presented a graph-based crossover for the STPG. PXST is based on partition crossover, firstly proposed to the TSP problem. The main adaptation is identifying partitions since the candidate solutions are tree-graphs, in this particular case.
Two soft-repair operators were applied in the GA using a graph representation of the individuals. The first repair relates to the property that the STPG can be solved by determining the MST of an appropriate set of Steiner vertices. An MST transformation calculates the MST spanning the vertices for a given ST. In this case, many edges can be replaced by lower-cost edges. Then, the GA would not necessarily explore all possible ST.
A second repair operation pruned non-terminal leaves, reducing the solution cost without losing the tree connectivity. The repair operator is applied since PXST and PrimRST might generate individuals with such leaves. In this first study, we do not analyze the impact of varying the numeric parameters (mutation or repair rate, for example).
Compared with two other GA for the STPG, the one using the PXST was faster for almost all instances. It also reached the global optimum solution for all instance problems with a success rate above 30% in general, considering 50 executions of the experiment.
PXST and PrimRST are similar considering the offspring are constituted only with edges of their parents, and common edges are transmitted to the offspring, most of them in the PrimRST case. The differences are that PXST does not delete edges or nodes present in the parents and exchanges more significant structures such as paths or subtrees. Because of their similarities, both crossover operators are at least competitive in their results.
The partition crossover operators, which PXST was based on, have the property of tunneling between optima [13]. It produces the best of \(2^k\) potential offspring, where k is the number of feasible partitions. Thus, if two parents are local optima, the offspring will be a local optimum with a lower cost. Considering that, PXST also can be used combined with other heuristics as in [13].
Furthermore, the GA using PXST is greedy and quickly converges to local or global optima. A mechanism to control the population diversity might contribute to GA with PXST performance.
In the PXST, some partitions remain unfeasible for recombination. Further studies can improve the partition discovery or fusion procedures to find more opportunities for recombination and improve the operator’s efficiency.
References
Beasley, J.E.: OR-library: distributing test problems by electronic mail. J. Oper. Res. Soc. 41(11), 1069–1072 (1990)
Cormen, T.H., Leiserson, C.E., Rivest, R.L., Stein, C.: Algoritmos: Teoria e Prática. Editora Elsevier (2012)
Esbensen, H.: Computing near-optimal solutions to the Steiner problem in a graph using a genetic algorithm. Networks 26(4), 173–185 (1995). https://doi.org/10.1002/net.3230260403
Hakimi, S.L.: Steiner’s problem in graphs and its implications. Networks 1(2), 113–133 (1971). https://doi.org/10.1002/net.3230010203
Kapsalis, A., Rayward-smith, V.J., Smith, G.D.: Solving the graphical Steiner tree problem using genetic algorithms. J. Oper. Res. Soc. 44(4), 397–406 (1993)
Karp, R.M.: Reducibility among Combinatorial Problems, pp. 85–103. Springer, Boston (1972). https://doi.org/10.1007/978-1-4684-2001-2_9
Kou, L., Markowsky, G., Berman, L.: A fast algorithm for Steiner trees. Acta Inf. 15(2), 141–145 (1981). https://doi.org/10.1007/BF00288961
Kruskal, J.B.: On the shortest spanning subtree of a graph and the traveling salesman problem. Proc. Am. Math. Soc. 7(1), 48–50 (1956). http://www.jstor.org/stable/2033241
Ljubić, I.: Solving Steiner trees: recent advances, challenges, and perspectives. Networks 77(2), 177–204 (2021)
Prim, R.C.: Shortest connection networks and some generalizations. Bell Syst. Tech. J. 36(6), 1389–1401 (1957)
Raidl, G.R., Julstrom, B.A.: Edge sets: an effective evolutionary coding of spanning trees. IEEE Trans. Evol. Comput. 7(3), 225–239 (2003). https://doi.org/10.1109/TEVC.2002.807275
Tinós, R., Whitley, D., Ochoa, G.: Generalized Asymmetric Partition Crossover (GAPX) for the asymmetric TSP. In: Proceedings of the 2014 Annual Conference on Genetic and Evolutionary Computation, GECCO 2014, pp. 501–508. ACM, New York (2014)
Tinós, R., Whitley, D., Ochoa, G.: A new generalized partition crossover for the traveling salesman problem: tunneling between local optima. Evol. Comput. 28(2), 255–288 (2020). https://doi.org/10.1162/evco_a_00254, pMID: 30900928
Whitley, D.: Next generation genetic algorithms: a user’s guide and tutorial. In: Gendreau, M., Potvin, J.-Y. (eds.) Handbook of Metaheuristics. ISORMS, vol. 272, pp. 245–274. Springer, Cham (2019). https://doi.org/10.1007/978-3-319-91086-4_8
Whitley, D., Hains, D., Howe, A.: Tunneling between optima: partition crossover for the traveling salesman problem. In: Proceedings of the 11th Annual Conference on Genetic and Evolutionary Computation, GECCO 2009, pp. 915–922. ACM, New York (2009)
Whitley, D., Hains, D., Howe, A.: A hybrid genetic algorithm for the traveling salesman problem using generalized partition crossover. In: Schaefer, R., Cotta, C., Kołodziej, J., Rudolph, G. (eds.) PPSN 2010. LNCS, vol. 6238, pp. 566–575. Springer, Heidelberg (2010). https://doi.org/10.1007/978-3-642-15844-5_57
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
de Godoi, G.A., Tinós, R., Sanches, D.S. (2021). A Graph-Based Crossover and Soft-Repair Operators for the Steiner Tree Problem. In: Britto, A., Valdivia Delgado, K. (eds) Intelligent Systems. BRACIS 2021. Lecture Notes in Computer Science(), vol 13073. Springer, Cham. https://doi.org/10.1007/978-3-030-91702-9_8
Download citation
DOI: https://doi.org/10.1007/978-3-030-91702-9_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-91701-2
Online ISBN: 978-3-030-91702-9
eBook Packages: Computer ScienceComputer Science (R0)