Abstract
The de-identification of clinical notes is crucial for the reuse of electronic clinical data and is a common Named Entity Recognition (NER) task. Neural language models provide a great improvement in Natural Language Processing (NLP) tasks, such as NER, when they are integrated with neural network methods. This paper evaluates the use of current state-of-the-art deep learning methods (Bi-LSTM-CRF) in the task of identifying patient names in clinical notes, for de-identification purposes. We used two corpora and three language models to evaluate which combination delivers the best performance. In our experiments, the specific corpus for the de-identification of clinical notes and a contextualized embedding with word embeddings achieved the best result: an F-measure of 0.94.
This work was partially supported by Institute of Artificial Intelligence in Healthcare, Memed, Google Latin America Research Awards, and by FCT under the project UIDB/00057/2020 (Portugal).
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
With the growth of Artificial Intelligence, predictions based on Electronic Health Records (EHRs) are becoming increasingly viable. As EHRs consolidate information from all patient timelines, they may be processed as input for machine learning models to build Clinical Decision Support Systems (CDSS), to improve hospital workflows. Several studies show the potential of using EHR data for comorbidity index prediction [24], fall event detection [26], and bringing smart screening systems to pharmacy services [23], along with many other possibilities [13].
Removing patient-identifiable health information is a prerequisite to conduct research over clinical notes [5]. In the United States, the Health Insurance Portability and Accountability Act (HIPAA) listed 18 types of personally identifiable information (PII) [7]. De-identification is the process of finding and removing PII from electronic health records.
De-identification is a typical Named Entity Recognition (NER) problem, part of the Natural Language Processing (NLP) field. The current state-of-the-art approach for NER problems is Bi-LSTM-CRF (bidirectional long-short-term-memory conditional random fields) [2], evaluated over a standard CoNLL-2003 corpus [19].
In this paper, we evaluate the use of neural network topology of Bi-LSTM-CRF using two corpora of patient name recognition in Portuguese. One patient-name-recognition corpus was annotated with 5 thousand names in 15 thousand sentences. Besides, a standard NER corpus was used to evaluate overfitting. Our approach focuses on the identification of the best cutoff point of the de-identification training set.
The rest of this paper is organized as follows: Sect. 2 presents previous works on the de-identification of clinical notes. Section 3 describes the datasets, language models, and neural network topology. We present the results in Sect. 4. Finally, in Sect. 5, we conclude and present future work.
2 Related Work
The de-identification of clinical notes has become a common task in the health information research field since the adoption of electronic health records [14]. Datasets such as i2b2 [29] and CEGS N-GRID [28] enable shared tasks in this research field, only for the English language.
In a recent survey, Leevy et al. [12] performed a systematic review of de-identification tasks in clinical notes using Recurrent Neural Networks (RNNs) and Conditional Random Fields (CRFs). They found 12 papers that used RNNs, 12 others that used CRFs, and finally 5 types of research that combine both approaches. Also, the authors consider that overfitting may be an issue when customized de-identification datasets are used during model training.
Most researches found in the survey lack the use of contextualized embeddings for the proposed task. The only exception is the work by Lee et al. [11]. In this investigation, they evaluate the de-identification performance of Gated Recurrent Units (GRUs) versus LSTM topologies, with a final CRF layer, with three types of embeddings: word-level embeddings, character-level embeddings, and contextualized embeddings.
To the best of our knowledge, no previous studies address the detection of patient names based on contextualized embeddings and token classification for the Portuguese language. We also evaluate the overfitting problem employing a non-customized corpus used for entity recognition in the general domain. In the next section, we cover the dataset, the neural network, and the language models (embeddings) used in the experiments.
3 Materials and Methods
In this section, we present the corpora used in the experiments, the neural network topologies, and the language models used in the neural networks for the representation of words. Besides, we describe how we evaluate the de-identification task.
The data used in the experiments we conducted for this article came from a project developed with several hospitals. Ethical approval to use the hospitals datasets in this research was granted by the National Research Ethics Committee under the number 46652521.9.0000.5530.
3.1 Data Source
We used two sources of Named Entity Recognition (NER) corpora for the sequence labeling task. The corpora are listed below:
EHR-Names Corpus: This corpus was built by collecting clinical notes from the aforementioned hospital (HNSC). We randomly selected 2,500 clinical notes for each hospital, totaling 15,952 sentences. The annotation process was performed by two annotators in the Doccano annotation tool [16]. The annotated dataset includes 12,762 tokens; 4,999 of them were labeled as a named entity.
HAREM Corpora: The HAREM corpora was an initiative of the Linguateca consortium. The first HAREM corpus [21] was published in 2006 with a human-made gold standard of named entities. The second HAREM corpus [20] was made available in 2008 with basically the same purpose. However, being a more recent contribution, the gold standard for the second HAREM consists in a slightly better test case. The HAREM corpora classify the named entities into ten categories. Nevertheless, for the purposes of our comparison, only the “Person" category was taken into account. Considering the “Person" category, the number of named entities is 1,040 for the first HAREM and 2,035 for the second HAREM.
3.2 Neural Network
In this section, we present the neural networks used for the downstream task evaluated in this study. For the neural networks, we used the FLAIR framework [1], developed in PyTorchFootnote 1; it has all the features of parameter tuning for model regulation. All our experiments were performed in NVIDIA T4 GPU.
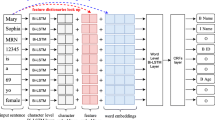
BiLSTM-CRF: As [2], we used a traditional sequence labeling task to learn how to detect names at the token level. Our sequence labeling is the product of training a BiLSTM neural network with a final CRF layer for token labeling. Bidirectional Long Short-Term Memory (BiLSTM) networks have achieved state-of-the-art results in NLP downstream tasks, mainly for sequential classifications [9, 17, 27].
Recurrent neural networks have proven to be an effective approach to language modeling, both in sequence labeling tasks such as part-of-speech tagging and in sequence classification tasks such as sentiment analysis and topic classification [10]. When it receives a sentence, this network topology concatenates information in both directions of the sentence. This makes the network have a larger context window, providing greater disambiguation of meanings and more accurate automatic feature extraction [8].
3.3 Language Models
WE-NILC: These are pre-computed language models that feature vectors generated from a large corpus of Brazilian Portuguese and European Portuguese, from varied sources and genres. Seventeen different corpora were used, totaling 1.3 billion tokens [6].
WE-EHR-Notes: For our experiments, we used a Word Embedding (WE) language model to vectorize the tokens of the clinical notes. We used a WE model that was previously trained by [22], where the authors reported the best results when using this WE trained with the neural network FastText [3]. The model was developed with 603 million tokens from clinical notes extracted from electronic medical records. The generated model has 300 dimensions per word. This model resulted in a vocabulary of 79,000 biomedical word vectors.
FlairBBP: This is a pre-trained model taught [25] with three corpora totaling 192 thousand sentences and 4.9 billion tokens in Portuguese. This language model was trained using Flair Embedding [2]. Flair is a contextual embedding model that takes into account the distribution of sequences of characters instead of only sequences of words, as is the case for Skip-gram and CBOW language models. Additionally, Flair Embeddings combine two models, one trained in a forward direction and one trained in a backward direction.
FlairBBP\(_{FnTg}\): This model was built using the aforementioned model FlairBBP fine-tuned by using the EHR-Names Corpus as input.
3.4 Design of the Experiments
Our evaluations were made within two sets of experiments: i) Stacking embeddings and ii) Corpus addition.
Experiments from set i aim to find the best combination of Language Models covered by this study. Therefore, we started by evaluating a BiLSTM-CRF neural network with shallow Word Embeddings only; then we employed FlairBBP, which is a contextualized language model; we stacked FlairBBP with a WE; and finally, we performed the experiments with the stacking of the fine-tuned FlairBBP with a WE.
Then, we performed set ii to evaluate the best cutoff point concerning the size of the training corpus of the model. Our strategy was to take several pieces of the corpus and train a model with each of these pieces in a cumulative sequence. In other words, we did these experiments by adding a corpus. This means that we divided the Corpus EHR-Names into 18 parts with 709 sentences each. Additionally, we added three more parts that are formed by the union of the EHR-Names corpus and one part of the HAREM corpus. Each part of HAREM also has 709 sentences. The corpora were added cumulatively, that is, the first split has 709 sentences, the second has 1,418, the third, 2.836, and so on. Each split corresponds to a trained model to find out which are the best metrics concerning the size of the corpus. Table 1 shows details of the sizes.
4 Results
Experiments from set i are shown in Table 2. In all the experiments in set i, we used Word2Vec Skip-Gram (W2V-SKPG) [15] Word Embeddings. Experiments from set ii are summarized in a plot in the image 1. Below we present our results and their interpretations from the comparison of the F1 metrics.
With the first set of experiments, we can see the impact of contextualized language models: the contextualized model FlairBBP outperforms WE-NILC by
 and WE-EHR-Notes by
and WE-EHR-Notes by
 . Another important aspect of the Contextualized Language Models is the potential combination with a Shallow WE, forming a final embedding with a high representational power of the language. Therefore, we managed to overcome both language models alone and we arrived at the best result of \(F_1 = 0.9413\), using FlairBBP stacked with WE-NILC. Our best result and consequently our best combination of embeddings were also found in [22, 25].
. Another important aspect of the Contextualized Language Models is the potential combination with a Shallow WE, forming a final embedding with a high representational power of the language. Therefore, we managed to overcome both language models alone and we arrived at the best result of \(F_1 = 0.9413\), using FlairBBP stacked with WE-NILC. Our best result and consequently our best combination of embeddings were also found in [22, 25].
In addition, our results with the stacks using FlairBBP\(_{FnTg}\) performed close to the average of the four stacks and both were slightly better than the stack of FlairBBP + WE-EHR-Notes.
From the second set of experiments, in which we looked for the best number of sentences for training, we found that the metrics PRECISION, RECALL, and F1 tend to grow up to split nine. After this split, there is a greater oscillation, as we can see in the graph of Fig. 1. Analyzing all the splits, we can see that split nine is the best, as it demands fewer corpus resources, naturally reduces the effort of the manual annotation of texts, and also reduces the computational cost, since the corpus is smaller.
Regarding the last three splits—which use the HAREM corpus—we can see that there is a significant drop in the metrics. We understand that the main reason for this is the difference in textual style between a medical record and the HAREM texts, extracted from magazines, newspapers, and interviews, among other genres, which are not related to health.
5 Conclusion
This paper presents an experimental study on the de-identification of patient names in electronic medical records. For the anonymization process, we manually developed a corpus in Portuguese with around 2,526 named entities and added the traditional HAREM corpus to our experiments. We used a BiLSTM-CRF neural network to train a de-identifier model and concluded that using Contextualized Language Models stacked with Word Embedding models produces cutting-edge results.
Additionally, our results show that it is not always necessary to have the entire training corpus to obtain the best predictor model and also that mixing very different textual genres can hinder the performance of the model. As future work, we plan to run experiments in generating language models, such as T5 [18] and GPT-3 [4].
All the content of the work (algorithm, sample dataset, language models, and experiments) is available at the project’s GitHub PageFootnote 2 in order to be easily replicated.
References
Akbik, A., Bergmann, T., Blythe, D., Rasul, K., Schweter, S., Vollgraf, R.: FLAIR: an easy-to-use framework for state-of-the-art NLP. In: Proceedings of the Conference of the North American Chapter of the Association for Computational Linguistics, pp. 54–59. Association for Computational Linguistics, Minneapolis, Minnesota, June 2019. https://doi.org/10.18653/v1/N19-4010, https://www.aclweb.org/anthology/N19-4010
Akbik, A., Blythe, D., Vollgraf, R.: Contextual string embeddings for sequence labeling. In: Proceedings of the 27th International Conference on Computational Linguistics, pp. 1638–1649 (2018)
Bojanowski, P., Grave, E., Joulin, A., Mikolov, T.: Enriching word vectors with subword information. Trans. Assoc. Comput. Linguist. 5, 135–146 (2017)
Brown, T.B., et al.: Language models are few-shot learners. In: Larochelle, H., Ranzato, M., Hadsell, R., Balcan, M., Lin, H. (eds.) Proceedings of the 33th Annual Conference on Neural Information Processing Systems (2020)
El Emam, K.: Guide to the De-identification of Personal Health Information. CRC Press, Boca Raton (2013)
Hartmann, N., Fonseca, E., Shulby, C., Treviso, M., Silva, J., Aluísio, S.: Portuguese word embeddings: evaluating on word analogies and natural language tasks. In: Proceedings of the 11th Brazilian Symposium in Information and Human Language Technology, pp. 122–131 (2017)
Hash, J., Bowen, P., Johnson, A., Smith, C., Steinberg, D.: An introductory resource guide for implementing the health insurance portability and accountability act (HIPAA) security rule. US Department of Commerce, Technology Administration, National Institute of \(\ldots \) (2005)
Hochreiter, S., Schmidhuber, J.: Long short-term memory. Neural Comput. 9(8), 1735–1780 (1997)
Jiang, Y., Hu, C., Xiao, T., Zhang, C., Zhu, J.: Improved differentiable architecture search for language modeling and named entity recognition. In: Proceedings of the 2019 Conference on Empirical Methods in Natural Language Processing, pp. 3585–3590. Association for Computational Linguistics, Hong Kong, China (2019)
Jurafsky, D., Martin, J.H.: Speech and Language Processing, vol. 3. Pearson, London, United Kingdom (2014)
Lee, K., Filannino, M., Uzuner, Ö.: An empirical test of GRUs and deep contextualized word representations on de-identification. In: MedInfo, pp. 218–222 (2019)
Leevy, J.L., Khoshgoftaar, T.M., Villanustre, F.: Survey on RNN and CRF models for de-identification of medical free text. J. Big Data 7(1), 1–22 (2020)
Magboo, Ma. Sheila A.., Coronel, Andrei D..: Data mining electronic health records to support evidence-based clinical decisions. In: Chen, Yen-Wei., Zimmermann, Alfred, Howlett, Robert J.., Jain, Lakhmi C.. (eds.) Innovation in Medicine and Healthcare Systems, and Multimedia. SIST, vol. 145, pp. 223–232. Springer, Singapore (2019). https://doi.org/10.1007/978-981-13-8566-7_22
Meystre, S.M., Friedlin, F.J., South, B.R., Shen, S., Samore, M.H.: Automatic de-identification of textual documents in the electronic health record: a review of recent research. BMC Med. Res. Methodol. 10(1), 1–16 (2010)
Mikolov, T., Chen, K., Corrado, G., Dean, J.: Efficient estimation of word representations in vector space. In: Bengio, Y., LeCun, Y. (eds.) Proceedings of the 1st International Conference on Learning Representations (2013)
Nakayama, H., Kubo, T., Kamura, J., Taniguchi, Y., Liang, X.: doccano: text annotation tool for human (2018). software available from https://github.com/doccano/doccano
Peters, M.E., et al.: Deep contextualized word representations. In: Proceedings of the Conference of the North American chapter of the association for computational linguistics: human language technologies, pp. 2227–2237 (2018)
Raffel, C., et al.: Exploring the limits of transfer learning with a unified text-to-text transformer. J. Mach. Learn. Res. 21, 140:1–140:67 (2020)
Sang, E.F., De Meulder, F.: Introduction to the CoNLL-2003 shared task: language-independent named entity recognition. arXiv preprint cs/0306050 (2003)
Santos, D., Freitas, C., Oliveira, H.G., Carvalho, P.: Second harem: new challenges and old wisdom. In: International Conference on Computational Processing of the Portuguese Language. pp. 212–215. Springer (2008). https://doi.org/10.1007/978-3-540-85980-2_22
Santos, D., Seco, N., Cardoso, N., Vilela, R.: Harem: An advanced NER evaluation contest for Portuguese. In: quot; In: Calzolari, N., et al. (ed.) Proceedings of the 5th International Conference on Language Resources and Evaluation (LREC 2006), Genoa Italy 22–28 May 2006 (2006)
dos Santos, H.D.P., Silva, A.P., Maciel, M.C.O., Burin, H.M.V., Urbanetto, J.S., Vieira, R.: Fall detection in EHR using word embeddings and deep learning. In: 2019 IEEE 19th International Conference on Bioinformatics and Bioengineering (BIBE), pp. 265–268, October 2019. https://doi.org/10.1109/BIBE.2019.00054
dos Santos, H.D.P., Ulbrich, A.H.D., Woloszyn, V., Vieira, R.: DDC-outlier: preventing medication errors using unsupervised learning. IEEE J. Biomed. Health Inform. 23, 8 (2018)
dos Santos, H.D.P., Ulbrich, A.H.D., Woloszyn, V., Vieira, R.: An initial investigation of the Charlson comorbidity index regression based on clinical notes. In: 2018 IEEE 31st International Symposium on Computer-Based Medical Systems (CBMS), pp. 6–11. IEEE (2018)
Santos, J., Consoli, B.S., dos Santos, C.N., Terra, J., Collovini, S., Vieira, R.: Assessing the impact of contextual embeddings for Portuguese named entity recognition. In: Proceedings of the 8th Brazilian Conference on Intelligent Systems, pp. 437–442 (2019)
Santos, J., dos Santos, H.D., Vieira, R.: Fall detection in clinical notes using language models and token classifier. In: 2020 IEEE 33rd International Symposium on Computer-Based Medical Systems (CBMS), pp. 283–288. IEEE (2020)
Straková, J., Straka, M., Hajic, J.: Neural architectures for nested NER through linearization. In: Proceedings of the 57th Annual Meeting of the Association for Computational Linguistics, pp. 5326–5331. Association for Computational Linguistics (2019)
Stubbs, A., Filannino, M., Uzuner, Ö.: De-identification of psychiatric intake records: overview of 2016 CEGS N-GRID shared tasks track 1. J. Biomed. Inform. 75, S4–S18 (2017)
Stubbs, A., Uzuner, Ö.: Annotating longitudinal clinical narratives for de-identification: The 2014 i2b2/UThealth corpus. J. Biomed. Inform. 58, S20–S29 (2015)
Acknowledgments
We thank Dr. Ana Helena D. P. S. Ulbrich, who provided the clinical notes dataset from the hospital, for her valuable cooperation. We also thank the volunteers of the Institute of Artificial Intelligence in Healthcare Celso Pereira and Ana Lúcia Dias, for the dataset annotation.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2021 Springer Nature Switzerland AG
About this paper
Cite this paper
Santos, J., dos Santos, H.D.P., Tabalipa, F., Vieira, R. (2021). De-Identification of Clinical Notes Using Contextualized Language Models and a Token Classifier. In: Britto, A., Valdivia Delgado, K. (eds) Intelligent Systems. BRACIS 2021. Lecture Notes in Computer Science(), vol 13074. Springer, Cham. https://doi.org/10.1007/978-3-030-91699-2_3
Download citation
DOI: https://doi.org/10.1007/978-3-030-91699-2_3
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-91698-5
Online ISBN: 978-3-030-91699-2
eBook Packages: Computer ScienceComputer Science (R0)