Abstract
Protein structure allows for an understanding of its function and enables the evaluation of possible interactions with other proteins. The molecular distance geometry problem (MDGP) regards determining a molecule’s three-dimensional (3D) structure based on the known distances between some pairs of atoms. An important application consists in finding 3D protein arrangements through data obtained by nuclear magnetic resonance (NMR). This work presents a study concerning the discretized version of the MDGP and the viability of employing genetic algorithms (GAs) to look for optimal solutions. We present computational results for input instances whose sizes varied from 10 to \(10^3\) atoms. The results obtained show that approaches to solving the discrete version of the MDGP based on GAs are promising.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
An important application in molecular biology is to find 3D arrangements of proteins using the interatomic distances obtained through nuclear magnetic resonance (NMR) experiments [24]. The objective of the Molecular Distance Geometry Problem (MDGP) is to determine molecular structures from a set of interatomic distances. Proteins are essential parts of living organisms. The interest in the 3D structure of proteins results from the intimate relationship between form and protein functions. The function of a molecule is determined by its chemical and geometrical structure [4]. Calculating the 3D structure of proteins is an essential problem for the pharmaceutical industry [16].
The MDGP was studied in [11] where a discretizable version of the MDGP (DMDGP) with a finite search space is also presented. The work proposes a Branch-and-Prune algorithm by searching a binary tree (representing the search space of the DMDGP) for a feasible solution to the problem. The MDGP can be formulated as an optimization problem whose objective is to determine, according to some criteria, the best alternative in a specified universe. In general, combinatorial optimization problems can be defined as given a set of candidate solutions S, a set of constraints \(\varOmega \), and an objective function f responsible for associating a cost f(s) to each candidate solution \(s \in S\). The objective is to find \(s^* \in S\) that is globally optimal and with the best cost amongst the candidate solutions [22]. Optimization methods can be divided into two classes: (i) deterministic, and (ii) stochastic. In the former, it is possible to predict the trajectory according to the input, i.e., the same input always leads to the same exit. Stochastic methods rely on randomness, thus the same input data can produce distinct outputs, as is the case with bio-inspired algorithms [9].
This work presents an evolutionary approach for solving the DMDGP and studies its respective viability. The DMDGP belongs to the class of NP-Hard problems through a reduction of the subset-sum problem [14]. Our aim is to answer the following questions: (i) how effective is an evolutionary heuristic for solving the DMDGP, evaluated on real 3D instances of proteins?; (ii) for which instance sizes is it possible to find optimal solutions? In order to tackle these issues we present two stochastic approaches based on genetic algorithms (GA) for solving the DMDGP.
The work is organized as follows. Related works are presented in Sect. 2. Section 3 formalizes the problem. The solution approaches proposals are characterized in Sect. 4. The computational experiments and approach comparisons are presented in Sect. 5. The concluding remarks are presented in Sect. 6.
2 Related Work
There are two ways to model the MDGP, namely, a continuous and a discrete formulation. There also exist disparities concerning the instances employed. Some works make use of geometric instances that were randomly generated based on a model [17]. Others employ synthetic instances produced in a random fashion based on a physical model [13]. There are also works that use dense real instances [18] whilst others employ sparse real instances [11] that were taken from the Protein Data Bank (PDB) [1].
Regarding the continuous version of the MDGP, in [6] the authors considered the case where the exact distances between all pairs of atoms are known. The algorithm is based on simple geometrical relationships between atom coordinates and the distances between them. The method was tested with the protein set HIV-1 RT that is composed of 4200 atoms. The algorithm is executed in linear time. In [12] a Divide and Conquer (D &C) algorithm was proposed that makes use of SDP alongside gradient descent refining. A structure consisting of 13000 atoms was solved in an hour. A geometric algorithm for approximate solutions with instances ranging from 402 to 7398 atoms was presented in [25].
In [5] the 3D-As-Synchronized-As-Possible (3D-ASAP) D &C approach was proposed. Protein lengths varied between 200 to 1009 atoms. The work also proposes a more agile version of 3D-ASAP, 3D-SP-ASAP, that uses a spectral partitioning algorithm to preprocess the graph into subgraphs. The results obtained showed that both are robust algorithms for sparse instances. The continuous approach most resembling this work was presented in [23] and describes the MemHPG memetic algorithm (MA). The method combines swarm intelligence alongside evolutionary computation associated with a local search with the objective of optimizing the crossover operator. MemHPG was tested with proteins whose lengths varied between 402 and 1003. The algorithm was able to obtain the solution to the problem. The result obtained was comparable with the then state of the art described in [25].
Regarding the discrete approaches, in [10], a D &C approach (ABBIE) was presented that explores combinatorial characteristics of the MDGP. The idea is to reduce the global optimization problem into subproblems. ABBIE was able to find approximate solutions for the MDGP. In [2] a distributed algorithm was described that does not make use of “known” positions to determine the successor ones. The method divides the problem graph into subgraphs using sampling methods: (i) SDP relaxation vying to minimize the sum of errors for specified and estimated distances; and (ii) gradient descent method of error reduction.
In the discrete formulations, the works most closely related to the one here presented are [21]. The authors define a discretizable variation of the Geometry Distance Problem known as the DMDGP and propose the Branch-and-Prune algorithm. In [8] a method is described to explore the symmetries that are present in the search space of the DMDGP and that can be represented by a binary tree. The method was able to obtain a relevant speedup for sparse instances against the Branch-and-Prune. Work [26] discusses a sequence of nested and overlapping DMDGP sub-problems. The results exhibited superior performance for randomly generated sparse instances.
3 Presentation and Modelling
3.1 DGP
The Distance Geometry Problem (DGP) consists in finding a set of points in a given geometric space of dimension k by knowing only some distances between them [15]. The DGP can be solved in linear time when all the distances are known [6]. Formally, given a simple, connected and weighted graph, \(G = (V, E, d)\), with a set V of n vertices, a set E of m edges, and a weight function \(d:E\rightarrow \mathbb {R}_+\) we wish to determine a function \(x: V \rightarrow \mathbb {R}^k\) such that \(||x(u) - x(v)|| = d(u, v)\) for every \(\{u, v\} \in E\). The DGP can be formulated as a global optimization problem whose objective function is described in Eq. (1).
where x solves the problem if and only if \(g(x) = 0\). This approach is challenging since an elevated number of local minima exist due to the non-convergence of the associated optimization problem.
3.2 MDGP
The 3D structure of a molecule is intrinsic to its function. Data provided by NMR allows for the calculation of the 3D structure of a protein. This procedure only provides the distances between close atoms. The problem consists in employing this information to obtain the position of every atom in the molecule. The MDGP is a particular case of DGP where the set of vertices V corresponds to the set of atoms of a molecule, the edges E is the set of known distances between atoms and the weight function d is the distance value.
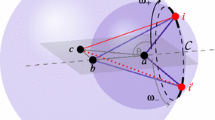
Figure 1(a) shows a solution in \(\mathbb {R}^2\) for a graph representing a problem instance. Atoms a, b, c can be fixed in \(\mathbb {R}^2\) to eliminate solutions from rotations and translations. Point d can be positioned in any of the countless points of the circle of center c and radius represented by the distance between c and d (Fig. 1(b)). The represented problem has a non-enumerable quantity of solutions. If the distance between b and d is also known (in addition to the one between c and d) then the problem presents a finite number of solutions. In this case, two solutions can be found at the intersection of the circumferences of center c and b with respective radius represented by the distances between: (i) c and d; and (ii) b and d. This case is exemplified in Fig. 1(c).
3.3 Discrete Version of the MDGP
As seen, the MDGP is a constraint satisfaction problem, which is generally reformulated as a global optimization one, where the objective is to minimize a penalty function capable of measuring how much the constraints are being violated (Eq. (1)). However, the penalty function of the optimization problem is strongly non-smooth. Optimization methods risk getting stuck at local minima with an objective value very close to the optimal one.
The DMDGP [19] is a particular case of MDGP when the geometric space is \(\mathbb {R}^3\), the set of vertices V is finite, and the set of edges E satisfies the following constraints: (i) every subset \(V_i=\{v_i, v_{i-1}, v_{i-2},v_{i-3}\}\subset V\) induces a \(4-\)clique in G; (ii) the distances in \(\{d(v_{i}, v_{j}):i,j\in V_i\}\) define non-null volume tetrahedrons. Therefore, when the discretization assumptions are satisfied, the domain of the penalty function can be reduced to a tree. In [11] the Branch-and-Prune method was proposed where the DMDGP search space is represented by a binary tree and the procedure attempts to fix an atom with each iteration. Assume that for each atom \(v_i\), for \(i > 3\), the distances between \(v_i\) and the tree immediately predecessors atoms \(v_{i-1}, v_{i-2}, v_{i-3}\) are known. Assuming that these three atoms possess realizations \(x_{i-1}\), \(x_{i-2}\) e \(x_{i-3}\) in \(\mathbb {R}^3\), then it is possible to define tree spheres centered in these points and whose radius is, respectively, the distances \(d(v_i,v_{i-1})\), \(d(v_i,v_{i-2})\) and \(d(v_i,v_{i-3})\). Under reasonable assumptions, the intersections of these spheres are the set \(\{x_i, x^\prime _i\}\) formed by the coordinates (branch) satisfying those three distances. One of these realizations can be chosen for vertex \(v_i\) and it is then possible to proceed to fix the next vertex \(v_{i+1}\). It may be possible that other edges (distances) exist that use vertex \(v_i\) that invalidate these coordinates. If this occurs then it becomes necessary to change the choices previously made (prune). This methodology defines a binary tree of atomic coordinates that contains every possible position for each vertex \(v_i\), \(i>3\), in the \(i^{\text {th}}\) tree layer. Each DMDGP solution is associated with a path from the root to a leaf node. This behaviour is exemplified in Fig. 2 where vector [0, 1, 1] represents a solution.
The representation of the DMDGP as a binary tree evidences several symmetry properties in the search space [20]. Namely, the invariance of the solution set concerning the total reflection of a solution is commonly referred to as the first symmetry. For example, if \(s = [0, 1, 1]\) in Fig. 2 is a feasible solution to the problem, then the inverse of all the bits in s (\(\overline{s} = [1, 0, 0]\)) is also a feasible solution. It is important to emphasize that, in the binary representation, the first three atoms are fixed in \(\mathbb {R}^3\). These characteristics led to the approach choice proposed in this work.
4 Solution Methodology
The solution approach developed to solve the DMDGP is based on the concepts of GAs. These are optimization algorithms imbued with a certain oriented randomness that is based on an analogy with the Darwinian principles of evolution and species genetics [7]. GAs have applications in discrete and continuous systems [27]. In this work, we propose two methods based on GAs, referred to as (i) Simplified Genetic Algorithm (SGA); (ii) Feasible Index Genetic Algorithm (FIGA). The main difference resides in the genetic operator employed. Each stage of the GAs proposed and their respective particularities are presented in the following subsections.
4.1 Representation and Initial Population
GAs are population algorithms where each individual represents a candidate solution for a problem. The population can be defined as a set of candidate solutions in the search space. The DMDGP search space is a binary tree, where each of the two possibilities in a given level of the tree symbolizes a position in \(\mathbb {R}^3\). A solution can be represented by a binary uni-dimensional chromosome of size n (number of atoms). The first three atoms have their position fixed. As a result, the chromosome can be implemented with size \(n-3\). The values assumed for each position, 0 or 1, are known as alleles and indicate the path chosen in the tree. The set of genes is known as the genotype. Figure 1 illustrates the coding implemented for the DMDGP. A constructive random heuristic generates the initial population. Namely, due to the coding of the problem for each position a choice is made (with 50% of probability) if the allele will have value 1 or 0.
4.2 Feasibility
The set of distances of the DMDGP can be divided into two distinct subsets, namely: (1) discretization distances; and (2) pruning distances. Discretization distances are the ones utilized in the construction of the binary tree, and pruning distances are the remaining ones. When only the subset of discretization distances is considered then all tree paths lead to feasible solutions. However, a vertex may exist in the tree that does not obey the pruning distances constraints. This results in the solution becoming unfeasible for that tree branch.
In the proposed FIGA algorithm, there is a method responsible for going through the tree and verifying until which chromosome position the solution is feasible, i.e., \(||x(u) - x(v)|| = d(u, v)\) for each \(\{u, v\} \in E\). This position, which will be referred to as the feasible index, is used to implement crossover and mutation operations. The SGA approach represents the basic implementation of a GA and therefore does not consider the feasible index in its genetic operators. This is the main difference between the previously proposed methods.
4.3 Fitness Function
At each generation, the individuals of a population are evaluated based on their fitness values. As is typical in optimization problems, the fitness function adopted in this work is the inverse of the objective function (OF) with 1 added in the denominator, as can be seen in Eq. (2).
The OF is equal to 0 when the known distances are equal to the calculated distances. In this case, x is a feasible solution for the DMDGP and the fitness will be equal to 1. The OF value is always greater or equal to zero. The fitness value tends to zero the bigger the OF value.
4.4 Selection
At each generation, the selection process elects the pairs of individuals that will participate in the crossover. These individuals are chosen based on a roulette selection that guarantees that all the individuals have a certain probability of being chosen that is proportional to their aptitude.
4.5 Crossover
Crossover is the process in which an offspring individual is created from the genetic information of the parents. This work employs a one-point crossover operator. In the SGA approach, the crossover consists in generating a random position between \([1, n-1]\), where n is the chromosome size, and the randomly selected position is the cut point of the SGA crossover. In the FIGA strategy, the crossover point is determined based on the feasible index (f) (Subsect. 4.2). This mechanism allows for the generated offspring to always inherit the feasible part of the parent that presents that largest feasible index. The FIGA crossover behavior is illustrated in Fig. 3. The feasible index for each parent is emphasized with the color red. The cut point is determined by the largest feasible index (dotted line). The interval position [0, c[ of the son, in which c is the cut point, will have the genetic information of the parent with the largest feasible index. The remaining values of the son (i.e., \(] c+1, n[\)) are copied from the other parent.
4.6 Mutation
The main objective of this operator is to introduce genetic diversity in the population of the current generation. Namely, each offspring generated can be mutated according to probability \(p_m\). If an offspring is selected for mutation, then the operator considers each gene individually and allows the allele to be inverted in accordance with the same probability. In the FIGA approach, the mutation is performed only on the infeasible part of the solution. Let f represent the feasible index (chromosome position in red - Fig. 4) and n the size of the chromosome, then the FIGA mutation can occur in the position of the interval \([f, n-1]\). In the SGA strategy, the mutation can happen in all the chromosome. Figure 4 illustrates the behavior of the FIGA mutation, with the feasible index being emphasized in red color.
4.7 Reset
The reset operation aims to avoid the solution getting stuck in the local minimum. Namely, if the fitness value of the best individual does not improve over a certain number of generations then the reset is activated. The best individual is maintained and the remaining population is discarded. A new population of individuals is generated with the random constructive heuristic.
4.8 Pseudocode
The pseudocode for the FIGA is presented in Algorithm 1. Both the SGA and the FIGA have as a stopping criterion a maximum quantity of generations (genMax) and search for a feasible solution (line 6). Line 1 initializes the variable that stores the largest fitness, bestFitness. The algorithm generates in a random fashion an initial population, calculates the fitness of each individual, and their respective feasible indexes (Lines 3, 5 and 5, respectively). The SGA does not consider the feasible index. Therefore, Line 5 is only present in the FIGA algorithm.
The second cycle (Line 8) considers the number of offspring formed with each generation. This value is defined by the multiplication of the crossover rate (\(t_c\)) and the population size (\(p_s\)). The line 9 selects a pair of individuals for the crossover operation. Line 10 is responsible for performing the crossover of the previously selected individuals. The newly generated offspring individual is stored in the variable offspring and can incur a mutation with probability \(p_m\). The mutation occurs in line 13. The line 15 (only available in FIGA) calculates the feasible index of the new individuals. As previously stated, the genetic operators (crossover and mutation) are different in the proposed algorithms. The fitness of the offspring is then compared against its respective parents. If an improvement is observed, the parent with worse fitness is replaced by its offspring (Lines 16 to 24). Both methods use the elitism scheme to prevent the loss of the fittest member of the population. Therefore, the method “getBestFiness” returns the largest fitness of the current population, this value is part of the stop criteria, and that individual will be part of the population for the next generation.
5 Results
5.1 Parameter Configuration
Due to the high-dimension search space, performing a full grid search of the optimization parameters would have a prohibitive cost. As a consequence, a smaller test was performed for synthetic instancesFootnote 1 in \(\mathbb {R}^2\). in order to define quality values for the GA parameters. The full set of results obtained is detailed in [3]. The evaluation procedure consisted of a set of instances with size \(n =\) [10, 30, 50, 100] was randomly generated taking into account the MDGP. These instances are composed of d lines, where d is the number of known distances. Each line represents a graph edge, i.e., a known distance between a pair of atoms. Each edge identifies the connecting atoms as well as the distance between them. For example, \([0 \ 1 \ 2.5]\) represents that atom \(v_0\) is at a distance of 2.5 measurement units from atom \(v_1\).
In these first tests, both methods were executed 10 times each in an algorithm independent run for each instance with each parameter combination. The parameters and values evaluated are presented in Table 2. There were 840 combinations of parameters totaling 8400 executions of the proposed algorithms for the four instance sizes considered in the study.
The tests allowed for a comparison between SGA and FIGA. For instances with only 10 atoms, it is not possible to observe which GA presents the best performance. However, as the number of atoms increases the balance shifts in favor of the FIGA. Namely, SGA is able to find a feasible solution for instances of small size (maximum 30 atoms). On the other hand, the FIGA is able to find the feasible solution for every instance size evaluated. By analyzing the results obtained the value were fixed in \(t_c = 0.7\), \(p_m = 0.3\), \(p_s = 200\) e \(genMax = 30\) values.
5.2 Experiments in 3D Space
This section presents the tests performed with real instances in \(\mathbb {R}^3\).
Instances.
The computational experiments described in this section employed real instances taken from the PDBFootnote 2 [1]. The instances selection was made randomly ensuring a linear increase in the number of atoms. The set of instances used is listed in Table 3 and illustrated in Fig. 5.
Presentation and Result Analysis. This section presents the FIGA results for the above mentioned \(\mathbb {R}^3\) instances. The elevated computational time made it unfeasible to perform a parameter study specific to the \(\mathbb {R}^3\) instances. As a consequence, the parameter values were empirically chosen based on the tests performed for \(\mathbb {R}^2\) and some tests for \(\mathbb {R}^3\) instances. After conducting empirical tests with various values for the reset, including not performing it, the value of 25 was selected for this parameter. Additional empirical tests were conducted considering values around the best parameters for \(\mathbb {R}^2\), as shown in Table 2. However, since the instances in \(\mathbb {R}^3\) are much larger than the synthetic instances in \(\mathbb {R}^2\), we had to increase the population size and the number of generations. Consequently, the parameter values for \(\mathbb {R}^3\) were defined as indicated in Table 4.
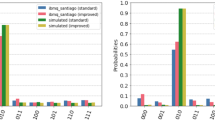
In the graphs that follow (Fig. 6), the green line depicts the largest fitness of the population, the blue line represents the average fitness and the red line corresponds to the smallest fitness.
Figure 6 shows the relationship between the feasible index and the number of generations for the instances with 114, 201, 300, 402, 501, and 702 atoms. When the algorithm finds a feasible solution the feasible index is equal to the chromosome size. The algorithm was able to find a feasible solution for all of the above mentioned instance sizes. However, the algorithm was unable to find a feasible solution for the remaining instances (Table 3 - 600, 801, 900, and 1002 atoms) using the set of stipulated parameters. By analyzing the results presented in the plots it is possible to gain a better understanding of the FIGA evolution until a feasible solution is found. For the smallest instance employed (Fig. 6(a)) the algorithm only required 6 generations to find a feasible solution. In the case of the instance using 702 atoms the algorithm was able to find a feasible solution after 350 generations. Table 5 indicates for each instance what was the iteration where a feasible solution was found or indicates with a character “–” when a solution was not found.
Future work would try to improve the GA in order to explore the symmetry properties of the DMDGP in the crossover and mutation operators. Potential improvement strategies: i) using a hierarchically structured population, ii) developing a memetic algorithm, and iii) evaluating other genetic operator types.
6 Final Considerations
This work presented a viability study regarding the application of GA for solving the DMDGP. The difference between the proposed algorithms, SGA and FIGA, resides in the partial or total feasibility of the solution and how this information affects the corresponding genetic operators. The FIGA evaluates a tree and verifies up until which chromosome position the solution is feasible. This index is used to perform crossover and mutation operations. The SGA does not consider this information in its genetic operators. Though subtle, this difference was responsible for significant improvements in the tests performed. For instances of size \(n =\) [10, 30, 50, 100] in \(\mathbb {R}^2\) the FIGA presented better or equal performance when compared against SGA. This performance was measured in terms of fitness and also the generation in which convergence occurred. The FIGA was able to solve real PDB instances for sizes 114, 201, 300, 402, 501, and 700.
Notes
- 1.
Instance set is publicly available in: https://bit.ly/3d0ezzo.
- 2.
The set of instances is available in https://www.rcsb.org/.
References
Berman, H.M., et al.: The protein data bank. Nucleic Acids Res. 28(1), 235–242 (2000)
Biswas, P., Toh, K.C., Ye, Y.: A distributed SDP approach for large-scale noisy anchor-free graph realization with applications to molecular conformation. SIAM J. Sci. Comput. 30(3), 1251–1277 (2008)
Carneiro, S., Souza, M., Filho, N., Tarrataca, L., Rosa, J., Assis, L.: Algoritmo genético aplicado ao problema da geometria de distâncias moleculares. In: Anais do LII Simpósio Brasileiro de Pesquisa Operacional (2020)
Creighton, T.E.: Proteins: Structures and Molecular Properties. Macmillan (1993)
Cucuringu, M., Singer, A., Cowburn, D.: Eigenvector synchronization, graph rigidity and the molecule problem. Inf. Infer. J. IMA 1(1), 21–67 (2012)
Dong, Q., Wu, Z.: A linear-time algorithm for solving the molecular distance geometry problem with exact inter-atomic distances. J. Global Optim. 22(1), 365–375 (2002)
Goldberg, D.E.: Genetic Algorithms in Search. Optimization, and Machine Learning (1989)
Goncalves, D.S., Lavor, C., Liberti, L., Souza, M.: A new algorithm for the \(^k\)dmdgp subclass of distance geometry problems (2020)
Gong, Y.J., et al.: Distributed evolutionary algorithms and their models: a survey of the state-of-the-art. Appl. Soft Comput. 34, 286–300 (2015)
Hendrickson, B.: The molecule problem: exploiting structure in global optimization. SIAM J. Optim. 5(4), 835–857 (1995)
Lavor, C., Liberti, L., Maculan, N., Mucherino, A.: The discretizable molecular distance geometry problem. Comput. Optim. Appl. 52(1), 115–146 (2012)
Leung, N.H.Z., Toh, K.C.: An SDP-based divide-and-conquer algorithm for large-scale noisy anchor-free graph realization. SIAM J. Sci. Comput. 31(6), 4351–4372 (2010)
Liberti, L., Lavor, C., Maculan, N., Marinelli, F.: Double variable neighbourhood search with smoothing for the molecular distance geometry problem. J. Global Optim. 43(2–3), 207–218 (2009)
Liberti, L., Lavor, C., Mucherino, A.: The discretizable molecular distance geometry problem seems easier on proteins. In: Mucherino, A., Lavor, C., Liberti, L., Maculan, N. (eds.) Distance Geometry, pp. 47–60. Springer, New York (2013). https://doi.org/10.1007/978-1-4614-5128-0_3
Liberti, L., Lavor, C., Mucherino, A., Maculan, N.: Molecular distance geometry methods: from continuous to discrete. Int. Trans. Oper. Res. 18(1), 33–51 (2011)
Maculan Filho, N., Lavor, C.C., de Souza, M.F., Alves, R.: Álgebra e Geometria no Cálculo de Estrutura Molecular. Colóquio Brasileiro de Matemática, IMPA (2017)
Moré, J.J., Wu, Z.: Global continuation for distance geometry problems. SIAM J. Optim. 7(3), 814–836 (1997)
Moré, J.J., Wu, Z.: Distance geometry optimization for protein structures. J. Global Optim. 15(3), 219–234 (1999)
Mucherino, A., Lavor, C., Liberti, L.: The discretizable distance geometry problem. Optim. Lett. 6(8), 1671–1686 (2012)
Mucherino, A., Lavor, C., Liberti, L.: Exploiting symmetry properties of the discretizable molecular distance geometry problem. J. Bioinform. Comput. Biol. 10(03), 1242009 (2012)
Mucherino, A., Liberti, L., Lavor, C.: MD-jeep: an implementation of a branch and prune algorithm for distance geometry problems. In: Fukuda, K., Hoeven, J., Joswig, M., Takayama, N. (eds.) ICMS 2010. LNCS, vol. 6327, pp. 186–197. Springer, Heidelberg (2010). https://doi.org/10.1007/978-3-642-15582-6_34
Mulati, M.H., Constantino, A.A., da Silva, A.F.: Otimização por colônia de formigas. In: Lopes, H.S., de Abreu Rodrigues, L.C., Steiner, M.T.A. (eds.) Meta-Heurísticas em Pesquisa Operacional, 1 edn., Chap. 4, pp. 53–68. Omnipax, Curitiba, PR (2013)
Nobile, M.S., Citrolo, A.G., Cazzaniga, P., Besozzi, D., Mauri, G.: A memetic hybrid method for the molecular distance geometry problem with incomplete information. In: 2014 IEEE Congress on Evolutionary Computation (CEC), pp. 1014–1021. IEEE (2014)
Schlick, T.: Molecular Modeling and Simulation: An Interdisciplinary Guide: An Interdisciplinary Guide, vol. 21. Springer, New York (2010). https://doi.org/10.1007/978-1-4419-6351-2
Sit, A., Wu, Z.: Solving a generalized distance geometry problem for protein structure determination. Bull. Math. Biol. 73(12), 2809–2836 (2011)
Souza, M., Gonçalves, D.S., Carvalho, L.M., Lavor, C., Liberti, L.: A new algorithm for a class of distance geometry problems. In: 18th Cologne-Twente Workshop on Graphs and Combinatorial Optimization (2020)
Yang, X.S.: Engineering Optimization: An Introduction with Metaheuristic Applications. Wiley (2010)
Acknowledgements
Douglas O. Cardoso acknowledges the financial support by the Foundation for Science and Technology (Fundação para a Ciência e a Tecnologia, FCT) through grant UIDB/05567/2020, and by the European Social Fund and programs Centro 2020 and Portugal 2020 through project CENTRO-04-3559-FSE-000158.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Carneiro, S.R.L., de Souza, M.F., Cardoso, D.O., Tarrataca, L., Assis, L.S. (2023). A Custom Bio-Inspired Algorithm for the Molecular Distance Geometry Problem. In: Naldi, M.C., Bianchi, R.A.C. (eds) Intelligent Systems. BRACIS 2023. Lecture Notes in Computer Science(), vol 14195. Springer, Cham. https://doi.org/10.1007/978-3-031-45368-7_12
Download citation
DOI: https://doi.org/10.1007/978-3-031-45368-7_12
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-45367-0
Online ISBN: 978-3-031-45368-7
eBook Packages: Computer ScienceComputer Science (R0)