Abstract
The critical scenario in public health triggered by COVID-19 intensified the demand for predictive models to assist in the diagnosis and prognosis of patients affected by this disease. This work evaluates several machine learning classifiers to predict the risk of COVID-19 mortality based on information available at the time of admission. We also apply a visualization technique based on a state-of-the-art explainability approach which, combined with a dimensionality reduction technique, allows drawing insights into the relationship between the features taken into account by the classifiers in their predictions. Our experiments on two real datasets showed promising results, reaching a sensitivity of up to 84% and an AUROC of 92% (95% CI, [0.89–0.95]).
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
The development of predictive models for the diagnosis and prognosis of patients has been recently investigated by numerous works [17]. The increasing interest in clinical predictive models may be explained by factors such as the growing capacity of data collection, the positive impact on decision-making and clinical care, and the great advances in machine learning (ML). These models aim to assist in the various challenges faced by health professionals while treating patients, optimizing clinical management processes, and allocating hospital resources, ultimately promoting better patient care.
The COVID-19 pandemic was a critical public health emergency worldwide, given the sudden need for many hospital beds and increased demand for medical equipment and health professionals. This scenario intensified the demand for resources and, consequently, the interest and need in using predictive models to support decision-making for diagnosis and prognosis in the context of COVID-19. As discussed by previous works [2], a considerable number of papers related to COVID-19 were published in a short period, including papers proposing ML applications for COVID-19 prognosis prediction.
Among the models for prognosis prediction, we can cite predictive models that aim to anticipate the patient’s outcome (e.g., hospital discharge or death), the severity of the disease, the hospitalization length, the evolution of the patient’s clinical condition during hospitalization, the need for intensive care unit (ICU) admission or interventions such as intubation, and the development of complications (e.g., cardiac problems, thrombosis, acute respiratory syndrome) [16, 17]. In this work, we are particularly interested in mortality prediction models.
Mortality prediction models help recognize patients with higher risks of poor outcomes at the time of admission or during hospitalization, providing support to more effective triage and treatment of ill patients and reducing death rates. Although previous studies have shown that ML algorithms achieved promising results in the mortality prediction task [2, 9, 18], the lack of explainability of the features taken into account by the predictive models has been pointed out as a limiting factor for their application in the clinical routine [9].
This work aims to evaluate several ML algorithms for predicting COVID-19 mortality and explore state-of-the-art explainability approaches to understand the factors considered during prediction. We developed a visualization technique based on SHAP values [11] combined with a dimensionality reduction method that draws insights into the relationship between the features taken into account by the predictive models and their correct and incorrect predictions. To assess our approach, we conducted a retrospective study of patients diagnosed with COVID-19, analyzing data collected during hospitalization in two hospitals. We also analyze the ability of classifiers to identify patients with a high probability of dying at different time intervals. Our main contributions can be summarized as follows: (i) evaluation of six ML algorithms for the mortality prediction task; (ii) proposal of a visualization technique based on SHAP values to assist in explainability; and (iii) experiments with real data from two hospitals.
Our findings showed that, in general, deceased patients were older. In addition, age, oxygen saturation, heart disease, and hemogram information also tended to have high contributions in the predictions. Logistic Regression, AdaBoost, and XGBoost models recurrently achieved the highest sensitivity scores among the evaluated classifiers.
2 Related Work
This section discusses existing works that addressed mortality prediction for COVID-19 patients based on the information available at hospital admission. We considered works evaluating classical risk scores (including those designed especially for COVID-19 patients and those that precede the pandemic) and ML classifiers. Several techniques have been employed to address mortality prediction, such as using risk scores based on classical statistical approaches and popular classification algorithms [5, 9, 20].
Covino et al. [5] evaluated six physiological scoring systems recurrently used as early warning scores for predicting the risk of hospitalization in an ICU and the risk of mortality—both tasks considering up to seven days. The Rapid Emergency Medicine Score obtained the best results with an AUROC of 0.823 (95% CI, [0.778–0.863]). Zhao et al. [20] developed specific risk scores for predicting mortality and ICU admission. The scores were developed based on logistic regression and reached an AUROC of 0.83 (95% CI, [0.73–0.92]). Another score specifically designed to predict mortality risk was the 4C Mortality Score [9]. It uses generalized additive models to categorize continuous feature values and pre-select predictors, followed by training a logistic regression model. The 4C score yielded an AUROC of 0.767 (95% CI, [0.760–0.773]), which was outperformed by the XGBoost algorithm by a small margin. Still, the authors claim that their score is preferable due to the difficulty in interpreting the predictions made by XGBoost. Paiva et al. [13] also evaluated classification algorithms for predicting mortality, including a transformer (FNet), convolutional neural networks (CNN), boosting algorithms (LightGBM), Random-Forest (RF), support vector machines (SVM), and K-Nearest-Neighbors (KNN). In addition, an ensemble meta-model was trained to combine all the aforementioned classifiers. The ensemble model achieved the best results, with an AUROC of 0.871 (95% CI [0.864–0.878]). To explain the predictive models, the study contemplated the application of SHapley Additive exPlanations (SHAP) et al. [11]. Araújo et al. [3] developed risk models to predict the probability of COVID-19 severity. To perform the study, the authors collected blood biomarkers data from COVID-19 patients. The models were developed with a LightGBM-based implementation, which achieved an average AUROC of 0.91 ± 0.01 for mortality prediction. The authors of this work apply a visualization technique similar to the one proposed in our work. The main difference is that they use it only to assess the model’s ability to separate the targeted classes. Conversely, in our work, we use visualization to analyze the relationship between the features taken into account by the classifiers and their correct and incorrect predictions.
The datasets used in the studies typically range from a single hospital unit with a few hundred patients to datasets comprising thousands of hospitalization records from hundreds of hospital units in several countries. Covino et al. [5] and Zhao et al. [20] carried out their studies based on datasets containing 334 and 641 patients, respectively, from a single hospital unit. Paiva et al. [13] used a dataset with 5,032 patients from 36 hospitals in 17 cities in 5 Brazilian states, and Araújo et al. [3] used a dataset with 6,979 patients from two Brazilian hospitals. To develop the 4C Mortality Score, Knight et al. [9] used a large dataset with 57,824 patients collected from 260 hospitals in England, Scotland, and Wales. Although the datasets used in the different studies do not contain the same features, there is a recurrence among the features considered most relevant by the predictive models. Age is the most frequently cited variable. Other features include oxygen saturation, respiratory rate, comorbidities, heart failure, lactate dehydrogenase (LDH), C-reactive protein (CRP), and consciousness level metrics (e.g., Glasgow Coma Scale).
Our study complements the aforementioned works by evaluating six algorithms in new datasets from two hospitals. In this work, in addition to applying the explainability approach, we developed a visualization technique that allows drawing insights into the relationship between the features taken into account by the classifiers and their correct and incorrect predictions.
3 Materials and Methods
This work aims to predict the risk of COVID-19 mortality during patient hospitalization based on information available at the time of admission. We approach this task through the use of ML classification algorithms.
Our method comprises five steps, depicted in Fig. 1. Our input dataset goes through preprocessing and cleaning, feature selection, then classification. The results of the classifiers are evaluated and submitted to the explainability approach. The next sections describe our data and each step in greater detail.
3.1 Patient Records and Preprocessing
The study involved patients hospitalized in two institutions: HMV and HCPA. HMV is a private hospital. HCPA is a public tertiary care teaching hospital academically linked to a University. The project was approved by the institutional Research Ethics Committee of both institutionsFootnote 1. Informed consent was waived due to the retrospective nature of the study. The collected data did not include any identifiable information to ensure patient privacy.
Patients included in the study are adults (age \(\ge \) 18) diagnosed with COVID-19 using the reverse transcriptase polymerase chain reaction (RT-PCR) test. The features include clinical and demographic data: age, sex, ethnicity, comorbidities (hypertension, obesity, diabetes, etc.), lab tests (arterial blood gas, lactate, blood count, platelets, C-reactive protein, d-dimer, creatine kinase (CK), creatinine, urea, sodium, potassium, bilirubin), need for ICU admission, type of ventilatory support during hospitalization and ICU period (O2 catheter, non-invasive ventilation, high-flow nasal cannula, invasive mechanical ventilation), start and end date of hospitalization, as well as hospitalization outcome. We performed the Shapiro-Wilk test on continuous features. The results showed that our features are not normally distributed (\(\alpha = 0.01\)).
Patients without information on the outcome of hospitalization (dependent variable) were not considered in this study. At the HCPA hospital, 318 patients had no record of outcome due to transfer to other hospitals and 6 due to withdrawal or evasion of treatment. For the HMV hospital, patients without outcome information were disregarded during data collection. Variables with missing values greater than 40% were also not considered. In the HMV dataset the variable brain natriuretic peptide was discarded, as well as the variables weight, height and blood type for HCPA. The variables troponin from HMV and fibrinogen from HCPA were the only exceptions to this rule, because although they had a missing value rate of 44.67% and 41.78%, respectively, there is previous evidence regarding their importance as indicators of severity for COVID-19. In HMV, 23 patients (5 deaths and 18 discharges) were disregarded because they had a missing value index greater than 30%, while in HCPA, 258 (93 deaths and 165 discharges) were not considered.
In HMV and HCPA, 7 and 19 patients, respectively, had two hospital admissions (readmissions). Among these, 7 patients from HMV and 13 patients from HCPA were readmitted within 30 days of the discharge from the index hospitalization. For these patients, we used the admission records collected at the index hospitalization with the readmission outcome as the label (patients who were discharged, returned within 30 days, and died, were considered deaths). For the other 6 patients from HCPA who were readmitted after 30 days, we used only the record of the index hospitalization.
HMV Datasets. contain the anonymized records of patients confirmed for COVID-19 admitted at HMV between March 2020 and June 2021. There are 1,526 records, among which 58.78% refer to male patients. The dataset covers both ward patients and patients admitted to the ICU. The data collection process was conducted by health professionals, who manually transcribed the information contained in the patients’ medical records into a form containing the features previously established for the study (clinical information available at hospital admission). Patients without outcome information were not considered during the data transcription process. Thus, our study did not include hospitalization records in which patients were transferred to other hospitals and/or dropout cases. Automatic verification processes were applied to identify inconsistencies. We developed classifiers considering two scenarios. In the first scenario, data of all patients are considered, including ward and ICU patients (\(\text {HMV}_{ALL}\)). In the second scenario, only patients admitted to the ICU are considered (\(\text {HMV}_{CTI}\)). In both cases, we focus on information available during admission.
\({\textbf {HCPA}}_{\boldsymbol{CTI}}\) Dataset contains anonymized records of 2,269 patients with confirmed COVID-19 admitted to the ICU of HCPA between March 2020 and August 2021. Male patients represent 54.69% of the dataset. The data was collected directly from the hospital database.
The outcome of interest was mortality during hospitalization, thus, records were categorized as alive or deceased according to data provided by the hospitals. Details of the datasets after the preprocessing step are shown in Table 1.
3.2 Feature Selection
Each raw instance was represented by roughly 60 and 70 features for HMV and HCPA datasets, respectively. Features with more than 40% of missing values were discarded. To pre-select features with good predictive power and avoid multicollinearity among predictive features, we applied the Correlation-based Feature Selection approach (CFS). This approach evaluates subsets of features prioritizing features highly correlated with the classes and uncorrelated with each other [7]. Feature selection was performed only over the training instances to avoid data leakage.
3.3 Classification
We applied six classification algorithms: (i) Logistic Regression (LR), three tree-based algorithms, namely (ii) J48, (iii) RandomForest (RF), and (iv) Gradient Boosting Tree - XGBoost (XGB), (v) a boosting-based ensemble (AdaBoostM1), and (vi) Multi Layer Perceptron (MLP).
In supervised classification approaches, class imbalance refers to the inequality of the proportions of instances for each class. Class imbalance is frequently observed in healthcare data, where the class of interest is often the minority (e.g., disease occurrence or poor diagnosis). When training classification algorithms on imbalanced data without the necessary care, these algorithms tend to present biased results, prioritizing the performance of the majority class. In our datasets, the outcome variable presents, to some degree, an imbalance between the classes (alive and deceased). The greatest imbalance is observed in the \(\text {HMV}_{ALL}\) dataset, with approximately 7.38 patients with an alive outcome for each patient with a deceased outcome. Considering only patients admitted to the ICU, the imbalance is 2.2 and 1.67 alive for each deceased, for the \(\text {HMV}_{CTI}\) and \(\text {HCPA}_{CTI}\) datasets, respectively.
To deal with class imbalance, we applied the Cost-sensitive matrix approach, in which the classification errors (false-negatives (FN) or false-positives (FP)) are weighted depending on the task to be addressed [14]. Since the \(outcome=deceased\) is our positive class, we applied a cost matrix that increased the penalty for FN. The cost matrices applied considered the imbalance between classes so that the penalty for false negatives was defined as \(pen = \frac{nc\_maj}{nc\_min}\) where \(nc\_maj\) is the number of instances of the majority class and \(nc\_min\) is the number of instances of the minority class.
3.4 Evaluation
Metrics. With the aim of evaluating the classification models, we calculated several metrics, namely: sensitivity (also known as recall or true positive rate), specificity (or true negative rate), positive predictive value (PPV or precision), negative predictive value (NPV), F1 (both for the positive class (F1\(_+\)) and macro-averaged (ma-F1) for both classes), Cohen’s kappa, Area Under Receiver Operating Characteristic Curve (AUROC), and Area Under Precision-Recall Curve (AUPRC). Using different metrics is important to interpret the results from different perspectives.
Resampling. We used the resampling technique Bootstrap to evaluate the predictive models. For each dataset, we performed 100 iterations of resampling with replacement. For each iteration, several instances—equal to the number of instances in the dataset—were randomly selected to compose the training set, and the remaining instances to constitute the test set. Thus, for each of the 100 iterations, the classifiers were trained on the training set and evaluated on the test set. At the end of the process, we take the means of the evaluation metrics.
3.5 Explainability of Classification Results
This section describes the proposed approach to help explain the features taken into account by the underlying model during inference. The goal is to extract insights into the relationship between the features utilized by the predictive models and their correct and incorrect predictions. The proposed approach corresponds to step four, shown in Fig. 1 and its components are shown in Fig. 2. The idea is similar to supervised clustering [10], in which SHAP values (not raw feature values) are used to cluster instances. However, instead of clustering, we use them to generate visual representations to inspect predictions.
First, starting from a generic dataset D, which in this case, could be any of our datasets, we use dimensionality reduction to understand aspects of linear separability of the data. It is expected that in the raw data, there are no clear separability patterns. Rectangles 2.a and 3.a represent the visualization of the raw data using a t-SNE projection [12]. We can see that the arrangement of instances does not constitute clusters, suggesting little linearity in the data.
Starting from the same dataset D, the proposed approach consists of the following steps: (2.b) training a classification algorithm and (3.b) applying the SHAP explanation over predictions. The training process in 2.b follows standard practices, such as handling missing values, applying techniques to deal with class imbalance, model evaluation, and calibration. In 3.b, we select instances from the test set and submit them to the SHAP approach. We obtain, for each feature value v of the instance i, their contribution values (SHAP values). In our experiments, we used KernelSHAP to estimate SHAP values, but other methods could also have been used (i.e., TreeSHAP, Linear SHAP [11], Kernel SHAP method to handle dependent features [1], etc.). With the SHAP values, we create an instance \(i'\), which consists of the calculated SHAP values instead of the raw feature values. In the next step (4.b), we create a dataset C constituted by the instances \(i'\).
Finally, dimensionality reduction is applied to the SHAP-transformed instances in C to generate a 2D plot that gives insights into the separability of instances. This procedure is represented by step 5.b. We applied the t-SNE dimensionality reduction (DR) technique in our tests, but other DR techniques could be used. The representation in 6.b is the result of the projection into 2D generated by t-SNE. We observe that the instances may form well-defined clusters in the projection. For instance, certain regions may have a prevalence of a specific class, while others may show higher uncertainty. Moreover, the projected points can be colored based on the feature values, which makes it possible to obtain important insights, such as the relationship between the features taken into account by the predictive models and their correct and incorrect predictions.
4 Experimental Evaluation
In this section, we describe the experiments performed to answer the following research questions:
- RQ1:
-
Is it possible to identify, among patients with COVID-19, who is more likely to die from the disease?
- RQ2:
-
Which features are taken into account by the classifiers for making predictions and how do they relate to correct/incorrect classifications?
- RQ3:
-
Is the performance obtained with multivariable classifiers superior to the performance obtained with single-variable classifiers using only age as independent variable?
4.1 Experimental Setup and Executions
Data preprocessing was performed using Python v3.5 and the libraries Pandas and Numpy. The classifiers were implemented using the Weka v3.9.6 API through the wrapper python-weka-wrapper3 v0.2.9Footnote 2—with the exception of the Gradient Boosting Tree, for which used the XGBoost implementationFootnote 3. XGBoost can handle missing values, so no data imputation was needed. For the other algorithms, the feature values were imputed with the median value from the training set. Categorical features were imputed with mode values. The weights of the cost matrix were calculated as shown in Sect. 3.3, considering the imbalance rate of the training set of each bootstrap iteration. To interpret the generated models, we use the SHAP libraryFootnote 4. SHAP relies on Shapley Values to explain the contribution that each variable makes to the output predicted by the model. Specifically, we used the KernelSHAP algorithm, as it is model-agnostic. The perplexity parameter for the t-SNE projection considered values between 10 and 40. Empirical tests were performed in multiple executions to set the appropriate perplexity value.
4.2 Results for Mortality Prediction (RQ1)
Table 2 displays the scores achieved by each classifier. Algorithms have better/worse performances across all metrics, with no clear winner. In our analysis, we give emphasis to sensitivity results, as the consequences of not identifying a patient with a high probability of dying are much more severe than wrongly predicting a patient into the alive class when a patient has a high probability of dying. Considering only sensitivity, Logistic Regression is the winner, leading all scenarios. Only in \(\text {HCPA}_{CTI}\) XGBoost reaches the same top score as LR. AdaBoostM1 has the second highest sensitivity values for \(\text {HMV}_{ALL}\) and \(\text {HMV}_{CTI}\), and the third highest sensitivity value for \(\text {HCPA}_{CTI}\). In all cases, J48 has the lowest sensitivity values.
Overall, the sensitivity values we obtain demonstrate the classifier’s promising ability to predict mortality based on the information available at the time of admission (answer to RQ1). The average sensitivity for the classification algorithms is higher for the \(\text {HMV}_{CTI}\) dataset (0.74) when compared to the \(\text {HCPA}_{CTI}\) dataset (0.65). This difference may be explained by the individual characteristics of patients admitted to each hospital and the features available in the datasets for each hospital. A similar discussion is mentioned in Futoma et al. [6], which warns about the risks of excessively seeking the development of predictive models that generalize to various hospitals with different clinical protocols and units.
Our best AUROC is higher than the ones reported in related work [5, 9, 13, 20] (see Table 2). Yet a direct comparison among approaches is not possible as they were applied to different datasets.
4.3 Analysis of Feature Importance and Explainability (RQ2)
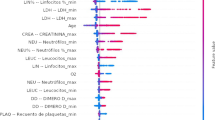
We use the results of applying dimensionality reduction over SHAP values to help understand the factors considered by classifiers during predictions and answer RQ2. Figure 3 presents the global view of how features contribute to mortality prediction using the LR classifier on \(\text {HMV}_{ALL}\). Each line refers to the contribution of a feature. Each dot is an instance. Low values for the feature are in blue and high values are in red. The contribution of the feature value is given by the horizontal displacement. The further the instance is positioned to the right, the greater the contribution to the deceased class. The further to the left, the greater the contribution of the feature value to the prediction of the alive class. For example, high values for age are associated with increased probability for the deceased class. The ten features with the highest contribution to the predictions were age, erythrocytes, bilirubin, oxygen saturation, neutrophils, troponin, heart disease, hemoglobin, creatinine, and central nervous system disease. Using the LR classifier on \(\text {HCPA}_{CTI}\), the ten features with the highest contribution were age, urea, red cell distribution width, D-dimer, mean corpuscular volume (MCV), lymphocytes, platelets, creatinine, erythrocytes, and troponin.
Figure 4 shows the t-SNE plots generated using SHAP values. Each point corresponds to a patient. The points are colored based on the values of different features to aid in interpreting the generated projections. Each row is composed of three plots. The first plot displays the correct and incorrect predictions (true positive (TP), true negative (TN), false positive (FP) and false negative(FN)). The second and third plots show points colored by different features.
The t-SNE plots reinforce the high importance attributed by the predictive models to the age feature. When analyzing the plots with the classifications and the colored plots based on the age of the patients (Figs. 4b, 4e 4h, and 4k), it is possible to observe that recurrent FP errors (purple dots) and TP occur in the regions with a high concentration of older patients. On the other hand, FN errors (red dots) and TN are in regions with younger patients.
Figure 4c also shows a trend of moderate and low erythrocyte values for TP and FP patients. These findings are supported by previous studies, which showed that anemia is associated with poor prognosis in critically ill patients.
Figures 4d, 4e, and 4f refer to LR on the \(\text {HMV}_{CTI}\) dataset. In Fig. 4f, we can observe that patients with heart disease tend to be older. Both features predispose those patients to be classified in the deceased class. This finding is in agreement with the related literature – studies have shown that heart disease is a risk factor for patient mortality [4].
Comparing the predictions for \(\text {HCPA}_{CTI}\) using XGB (Figs. 4g, 4h, and 4i) and LR (Figs. 4j, 4k, and 4l) we notice that XGB tends to present a greater overlap between FN and FP. This may be due to the influence of the t-SNE visualization method, which may favor the visualization for the LR algorithm.
Figures 4i and 4l, generated based on the SHAP values provided by the XGB and LR classifiers, show regions of high concentrations of patients with high values of D-dimers. Both younger and older patients are found in those regions. The pattern is more evident in the visualization obtained with LR, in which TP and FP classifications prevail. High D-dimer values are associated with poor prognosis in patients with COVID-19 infection [15], and in both cases (XGB and LR), models tend to assign these patients to the deceased class.
4.4 Single vs. Multiple Variable Classifiers (RQ3)
As seen in the analysis of Sect. 4.3, as well as recurrently pointed out by works in the related literature, age was frequently flagged as the attribute with the greatest contribution during mortality prediction tasks. This research question aims to compare the performance between classifiers with multiple independent variables and their respective versions with only one independent variable. For this, we compare the performance of the classifiers presented in Sect. 4.2. Figure 5 shows the differences between the performances obtained by the classifiers trained using only age as independent variable.
For most algorithms, classification models that used only age as independent variable did not show degradation in sensitivity, and in some cases, higher levels of sensitivity were found. Although this positive effect on sensitivity levels was observed, the reduction in PPV and specificity levels was also recurrent. As a consequence, in most cases, no gains were observed for the metrics F1+, ma-F1, Kappa, AUROC and AUPRC. The LR algorithm, which showed the best results in the experiments in Sect. 4.2, tended not to show substantial decrease in sensitivity levels. For the \(\text {HMV}_{ALL}\) dataset, considerable increases in sensitivity levels occur for some algorithms. In contrast, there were considerable reductions in the levels of PPV and specificity, mainly in the \(\text {HMV}_{ALL}\) dataset and in the \(\text {HCPA}_{CTI}\) dataset.
Algorithm J48, which presented the lowest sensitivity scores in the experiment in Sect. 4.2, was recurrently the one that obtained the highest sensitivity gains among the classifiers that used only age as an independent variable. When considering the \(\text {HMV}_{CTI}\) dataset, in addition to the J48 algorithm, the RF and MLP algorithms stood out for their increased sensitivity. However, despite the increase in sensitivity, due to the losses in the levels of PPV and specificity, no gains were observed for the metrics F1+, ma-F1 and Kappa. Finally, J48 and MLP were the only ones that obtained AUROC gains. In \(\text {HCPA}_{CTI}\) (Fig. 5b), only the J48 algorithm obtained an increase in the other metrics, in addition to sensitivity. In \(\text {HMV}_{CTI}\) (Fig. 5c), J48 and MLP achieved the highest sensitivity gains. In this scenario, both algorithms showed a smaller reduction in terms of PPV and specificity, resulting in an increase in the levels of other metrics.
Essentially, the classifiers that use only age as independent variable reach similar or higher sensitivity scores compared to their multivariable versions. However, these scores were achieved at the cost of lower Spe and PPV levels, showing that other variables are useful in improving these metrics. The full version of Table 2 with CI (RQ1) and significance tests performed on RQ3 can be accessed at https://github.com/danimtk/covid-19-paper.
5 Conclusion
This work evaluated several ML algorithms for predicting COVID-19 mortality. The classifiers showed a promising ability to predict mortality based on the information available at the time of admission, reaching a sensitivity of up to 84% and an AUROC of 92% (95% CI, [0.89–0.95]). Overall, besides age, oxygen saturation, heart disease, hemogram information also tended to have high contributions to the predictions. We applied a visualization technique based on SHAP values, which helps to corroborate the high importance attributed to the age feature by the classifiers in their predictions. The visualizations also showed that some groups of patients with low erythrocytes and high D-dimer (risk factors for mortality) tend to be recognized by the classifiers with a higher risk of dying. We believe that the visualization technique employed here can also be used in other prediction tasks.
One limitation of our work is that SHAP is relatively time-consuming when applied to datasets with a large number of features [19]. This limitation can be mitigated using optimized SHAP algorithms (like TreeSHAP or [19]). Although the visualizations obtained are intuitive and seem to represent already known relationships (such as the high risk for older patients, as well as those with comorbidities), it should be noted that approaches that use Shapley values, as well as approximations of these values (such as SHAP), may generate misconceptions regarding the actual contribution of features [8]. Thus, we emphasize the importance for specialists to use other evaluation techniques concomitantly to assist in the interpretation of predictive models, avoiding misinterpretations.
In future work, we plan to experiment with other dimensionality reduction methods and classifiers. We also plan to investigate whether there is a set of attributes (and how many) would be needed to maintain sensitivity levels without considerable PPV losses.
Notes
- 1.
Ethics committee approval: HCPA 32314720.8.0000.5327, HMV 32314720.8.3001.5330.
- 2.
- 3.
- 4.
References
Aas, K., Jullum, M., Løland, A.: Explaining individual predictions when features are dependent: accurate approximations to Shapley values. Artif. Intell. 298, 103502 (2021)
Alballa, N., Al-Turaiki, I.: Machine learning approaches in Covid-19 diagnosis, mortality, and severity risk prediction: a review. Inf. Med. Unlocked 24, 100564 (2021)
Araújo, D.C., Veloso, A.A., Borges, K.B.G., das Graças Carvalho, M.: Prognosing the risk of Covid-19 death through a machine learning-based routine blood panel: a retrospective study in brazil. IJMEDI 165, 104835 (2022)
Broberg, C.S., Kovacs, A.H., Sadeghi, S., et al.: Covid-19 in adults with congenital heart disease. JACC 77(13), 1644–1655 (2021)
Covino, M., Sandroni, C., Santoro, M., et al.: Predicting intensive care unit admission and death for Covid-19 patients in the emergency department using early warning scores. Resuscitation 156, 84–91 (2020)
Futoma, J., Simons, M., Panch, T., et al.: The myth of generalisability in clinical research and machine learning in health care. Lancet Digit. Health 2(9), 489–492 (2020)
Hall, M.A.: Correlation-based feature selection for machine learning. Ph.D. thesis, The University of Waikato (1999)
Huang, X., Marques-Silva, J.: The inadequacy of Shapley values for explainability. arXiv preprint arXiv:2302.08160 (2023)
Knight, S.R., Ho, A., Pius, R., et al.: Risk stratification of patients admitted to hospital with Covid-19 using the ISARIC WHO clinical characterisation protocol: development and validation of the 4C mortality score. BMJ 370, m3339 (2020)
Lundberg, S.M., Erion, G.G., Lee, S.I.: Consistent individualized feature attribution for tree ensembles. arXiv preprint arXiv:1802.03888 (2018)
Lundberg, S.M., Lee, S.I.: A unified approach to interpreting model predictions. In: Advances in Neural Information Processing Systems, vol. 30 (2017)
Van der Maaten, L., Hinton, G.: Visualizing data using t-SNE. JMLR 9(11), 2579–2605 (2008)
Miranda de Paiva, B.B., Delfino-Pereira, P., de Andrade, C.M.V., et al.: Effectiveness, explainability and reliability of machine meta-learning methods for predicting mortality in patients with COVID-19: results of the Brazilian COVID-19 registry. medRxiv (2021)
Qin, Z., Zhang, C., Wang, T., Zhang, S.: Cost sensitive classification in data mining. In: Advanced Data Mining and Applications, pp. 1–11 (2010)
Rostami, M., Mansouritorghabeh, H.: D-dimer level in COVID-19 infection: a systematic review. Exp. Rev. Hematol. 13(11), 1265–1275 (2020)
Subudhi, S., Verma, A., Patel, A.B.: Prognostic machine learning models for Covid-19 to facilitate decision making. IJCP 74(12), e13685 (2020)
Wynants, L., Van Calster, B., Collins, G.S., et al.: Prediction models for diagnosis and prognosis of COVID-19: systematic review and critical appraisal. bmj 369, m1328 (2020)
Yadaw, A.S., Li, Y., Bose, S., et al.: Clinical features of COVID-19 mortality: development and validation of a clinical prediction model. Lancet Digt. Health 2(10), E516–E525 (2020)
Yang, J.: Fast TreeSHAP: accelerating SHAP value computation for trees. arXiv preprint arXiv:2109.09847 (2021)
Zhao, Z., Chen, A., Hou, W., et al.: Prediction model and risk scores of ICU admission and mortality in COVID-19. PLoS ONE 15(7), e0236618 (2020)
Acknowledgment
This work has been financed in part by CAPES Finance Code 001 and CNPq/Brazil.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Kuhn, D.M., de Loreto, M.S., Recamonde-Mendoza, M., Comba, J.L.D., Moreira, V.P. (2023). Explainability of COVID-19 Classification Models Using Dimensionality Reduction of SHAP Values. In: Naldi, M.C., Bianchi, R.A.C. (eds) Intelligent Systems. BRACIS 2023. Lecture Notes in Computer Science(), vol 14195. Springer, Cham. https://doi.org/10.1007/978-3-031-45368-7_27
Download citation
DOI: https://doi.org/10.1007/978-3-031-45368-7_27
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-45367-0
Online ISBN: 978-3-031-45368-7
eBook Packages: Computer ScienceComputer Science (R0)