Abstract
The human gastrointestinal tract is prone to various abnormalities, including lethal diseases such as cancer, necessitating better endoscopic performance and standardized screening. Endoscopic scoring systems lack generalizability, emphasizing the need for artificial intelligence-based solutions. Using the HyperKvasir dataset, we employed deep learning, specifically Convolutional Neural Networks, or shortly CNNs, to analyze endoscopic images and videos. Our study focused on improving the classification of gastrointestinal tract diseases by proposing various CNN ensembles and fusion techniques. Through the use of seven CNN models and effective merging techniques, we achieved enhanced performance. Validation involved literature review and experiments. DenseNet-161 influenced the merger process, and integrating ResNet152 and VGG further enhanced effectiveness. Resource analysis included GPU model, RAM usage, and execution time. Results demonstrated comparable performance to the previous model, with F1-score of 0.910 and Matthews correlation coefficient, MCC for short, of 0.902, using 10 GB GPU RAM (compared to 15.8 GB). With 24.7 GB GPU RAM, F1-score of 0.913 and MCC of 0.905 were achieved. These findings advance our understanding of ensemble architectures and fusion techniques.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
The human gastrointestinal (GI) tract is susceptible to several abnormal mucosal findings, including life-threatening diseases [2]. GI cancer alone accounts for millions of new cases annually, emphasizing the need for improved endoscopic performance and systematic screening [10]. Gastrointestinal exams and colonoscopy are essential procedures to investigate the human GI tract [9]. These tests play a vital role in the diagnosis and management of gastrointestinal conditions, contributing to the early detection, treatment and prevention of serious complications [2, 5, 7]. However, current endoscopic scoring systems lack standardization and are subjective [2, 7].
In this context, artificial intelligence (AI)-enabled computer-assisted diagnostic systems, particularly machine learning, have shown promise in healthcare, but the scarcity of medical data impedes progress [2, 8]. To solve this, we used a database (DB), called HyperKvasir, a large dataset of gastrointestinal images and videos collected during real exams [2]. The dataset contains over \(1.1 \times 10^5\) images and 374 videos and represents anatomical landmarks as well as pathological and normal findings [2].
Over the years, machine learning has evolved into deep learning algorithms, relying primarily on the Deep Neural Network (DNN). Convolutional neural networks (CNN), a type of DNN, have emerged as a powerful tool for image analysis and classification, including medical imaging tasks. CNN Ensemble architectures have been widely employed to improve forecast accuracy by combining the outputs of various models. These sets leverage the diversity of individual CNN models to improve overall performance. In addition, fusion techniques are employed to effectively integrate predictions from multiple CNN models [11, 16].
In this work, our main objective is to propose a new ensemble architecture and efficient fusion techniques for CNNs in the classification of gastrointestinal tract diseases using the HyperKvasir dataset, aiming to obtain better results than in the literature and to optimize computational resources. To achieve this, we performed a thorough literature review to identify relevant studies on the use of deep learning methods in similar health domains. In addition, we performed several experiments to evaluate the effectiveness of our proposed approach.
2 Theoretical Foundation
Since the emergence of computers, there have been research efforts to make them reproduce biological characteristics; these are known as bioinspired research. Among the bioinspired research, there is one that seeks to simulate the functioning of the brain through artificial neural networks. These networks have undergone many transformations, as reported in the papers [4, 11,12,13, 16,17,18]. This section provides an overview of the HyperKvasir database [2], the dataset utilized in this study. We reviewed the literature on deep learning in digital imaging (DI) and consider Borgli et al.’s general model [2] as a reference for our research. Our objective is to establish a robust foundation by analyzing the dataset and surveying related studies.
2.1 HyperKvasir Dataset
The HyperKvasir datasetFootnote 1 is composed of images and videos. The dataset content was collected between 2008 and 2016, in a hospital in Norway. In this work, 10639 labels available in the dataset are used. The images are separated into 23 classes, in Fig. 1 it is possible to see examples of images contained in the dataset. Class labels are Z-line, Pylorus, Retroflex stomach, Barrett’s, Short segment, Esophagitis grade A, Esophagitis grade B-D, Cecum, Retroflex rectum, Terminal ileum, Polyps, Ulcerative colitis grade 0–1, Ulcerative colitis grade 1, Ulcerative colitis grade 1–2, Ulcerative colitis grade 2, Ulcerative colitis grade \(2-3\), Ulcerative colitis grade 3, Hemorrhoids, Dyed lifted polyps, Dyed resection margins, BBPS 0–1, BBPS 2–3, Impacted stool. The dataset offers a file (.csv) with the division of classes studied by Borgli [2].
2.2 Related Works
In our study, we started by establishing a solid foundation using the reference article [2]. Expanding upon this work, we created a comprehensive graph, as illustrated in Fig. 2, to visually illustrate the interconnectedness of relevant papers in our research field. This graph provides a valuable representation of the network of related literature, with a specific focus on the HyperKvasir image and video dataset for gastrointestinal endoscopy, as discussed by Borgli et al. [2].
Graph for connected papers with HyperKvasir image and video dataset for gastrointestinal endoscopy, Borgli et al. [2].
Upon analyzing the graph, we identified a total of 41 studies connected to the article [2], resulting in a set of 42 relevant studies for our research. However, we established inclusion criteria, considering only studies published after 2019, that is, after the publication date of the base article. Additionally, we excluded systematic literature reviews or survey studies from our analysis. The 10 remaining studies were evaluated for their degree of similarity to the base article, represented by the similarity index SbP (similar based-paper), ranging from 12% to 100%. The higher the SbPFootnote 2 value, the greater the similarity between the article and the base work (previous paper [2]), which is relevant to the results obtained in our research.
To gather related works for our paper, each article was thoroughly reviewed based on the following parameters: Study name and year, Task performed, CNN Architecture used, Methodology approach, Dataset and (SbP) Similarity on the base paper. These parameters were used to assess and categorize the papers, ensuring that they align with the focus and objectives of our research. By analyzing each article based on these criteria, we were able to identify and select relevant works that contribute to our study.
Table 1 shows several studies in the context of gastrointestinal endoscopy. The studies cover a range of tasks such as polyp classification, segmentation, detection, localization, and abnormality identification. Various deep learning architectures, including CNNs like ResNet-152, DenseNet-161, U-Net, Pix2Pix and HGANet are utilized in these studies. Different methodologies and techniques such as Fuzzy C-Means Clustering, ResUNet, Conditional Random Fields (CRF), Test-Time Augmentation (TTA), and Adversarial Training are also employed. Multiple datasets are used for evaluation, including the HyperKvasir and the EAD2019 datasets. The achieved similar based-paper (SbP) rank is used as a performance metric, with higher values indicating better results, which means is most similar to paper [2].
These studies provided valuable insights and advancements in leveraging deep learning and clustering techniques for gastrointestinal endoscopy. They contribute to the development of automated systems for polyp classification, segmentation, and abnormality detection, which can improve the efficiency and accuracy of medical diagnoses. Besides that, these papers served as the foundation for our study, as they utilized various CNNs for different approaches to GI problems.
2.3 Background Model
In Borgli et al. [2], the authors proposed a classification model represented in Fig. 3. The model is composed of pre-trained CNNs, DenseNet161 and ResNet152. Each CNN model is a function, M, composed of a set of subfunctions (convolution, pooling, batch normalization, softmax, optimize, etc.) which, in this case, given an input image \(\overrightarrow{\chi }\), a learning rate value \(\eta \) and the number of epochs e, returns an output \(\overrightarrow{P}\), according to the function:
Previous model [2] for CNN models training and fusion process.
The output, \(\overrightarrow{P}\), is a probability vector that indicates the probability that \(\overrightarrow{v}_i\) belongs to one of the classes of the problem. The vector \(\overrightarrow{P} =[c_1, c_2, c_3, \cdots , c_{23}]\), where \(c \in C\) for each GI class and \(|C| = 23\). Given a dataset with 10639 gastrointestinal images, \(\overrightarrow{\chi }\), divided into two splits, one set with 5315 images and another set with 5324 images. The model proposed in [2] alternates sets for training and validation. The final response of the model is the average of the results of the two splits. For each split, the authors trained the CNNs models \(M_{1}\) and \(M_{2}\), respectively, DenseNet161 and ResNet152, using \(\eta = 0.001\), \(e = 50\) and optimize SGD, stochastic gradient descent; \(M_{1}\) and \(M_{2}\) generated the responses \(\overrightarrow{P}_{1}\) and \(\overrightarrow{P}_{2}\), respectively. After training, the best weight set \(\overrightarrow{w}\) for each model was found and the best-trained models \(M_{1}^{b}\) and \(M_{2}^{b}\) are saved. Using the trained models, a model \(M^{\nu }\) is created:
since \(\overrightarrow{P}_{i}^{\nu } = \frac{(\overrightarrow{P}_{1}^{b} + \overrightarrow{P}_{2}^{b})}{2}\), \(\overrightarrow{P}_{1}^{b}\) and \(\overrightarrow{P}_{2}^{b}\) are output of \(M_{1}^{b}\) and \(M_{2}^{b}\), respectively. For each split, the CNN model \(M^{\nu }\) was trained using \(\eta = 0.001\) and \(e = 50\).
3 Proposal
In this paper, we have two proposals. The first proposal sought to overcome the literature results using fewer GPU resources than the second proposal, which is an expansion of what the authors propose in [2]. The second proposal sought to verify whether other CNN fusions, even using more GPU resources, outperformed the literature results. The CNNs models used are accessible through the Pytorch framework.
3.1 Fusion and Ensemble Processes
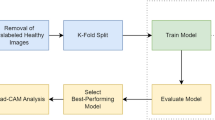
Our first proposal was to individually train a set of CNNs models \(\{M_{1}, M_{2},\) \(M_{3}, ..., M_{n}\}\) and get their respective answers \(\{\overrightarrow{P}_{1}, \overrightarrow{P}_{2}, \cdots , \overrightarrow{P}_{n}\}\), for \(n = 7\). The CNNs were DenseNet121, DenseNet161, DenseNet201, EfficientNet_b0, MobileNetV2, ResNet152 e VGG16. Each model was trained adopting the following values for \(\eta = \{0.0001,\) 0.0003, 0.0005, 0.001, 0.003, \(0.005\}\), \(e = 50\) and optimize SGD, as shown in Fig. 4. After training, each model was tested and the responses were fused. Each network could participate or not in the fusion. Fusions occur between trained models with the same learning rate, the total of fusions were \(2^{n} \times 6\).
For this proposal, two fusion alternatives were analyzed, by average and by voting. In average fusion, each trained model, \(M_{i}\), generates an output \(\overrightarrow{P}_{i}\) and the fusion is given by \(\overrightarrow{P}_{\tau } = \frac{1}{n}\sum _{i=1}^{n} \overrightarrow{P}_{i}\), where n is the number of models of CNNs and \(\overrightarrow{P}_{\tau }\) is the average output of the models that make up the fusion, according to Fig. 5(a).
In fusion by voting, considering the values in Fig. 5(a), each network that makes up the fusion votes in the class that receives the highest percentage of probability, as shown in Fig. 5(b). In case of a tie, the first tiebreaker considered the number of times the class was in first and second place in the voting. If the tie remains, among the classes that met the first tiebreaker criterion, the one with the highest percentage of probability is chosen.
In our second proposal, we created models with different fusions of pre-trained CNNs, so \(M^{\nu }\) was the composition of best models \(\{M_{1}^{b}\), \(M_{2}^{b}\), \(\cdots , M_{n}^{b}\}\), for different values of \(2 \le n \le 7\). The results of the proposals are presented in Sect. 4. In the first proposal, with trained models, it is possible to perform fusions between models without having to train them again. In the second proposal, each new fusion generates a new model that needs to be trained.
3.2 Evaluation Methodology
Our approach involved the utilization of four metrics - precision, recall, F1-score, and Matthews correlation coefficient (MCC) – to evaluate the performance of our model and gain valuable insights. Additionally, we employed both macro and micro averages to further analyze the overall performance of our model [15].
In classification tasks, true positive (TP), true negative (TN), false positive (FP), and false negative (FN) are commonly used terms that represent the outcomes of the predictions made by a model. TP refers to correct predictions of the positive class, where the model identifies positive instances correctly. TN represents correct predictions of the negative class, where the model identifies negative instances correctly. FP refers to incorrect predictions of the positive class, where the model identifies negative instances as positive. FN represents incorrect predictions of the negative class, where the model identifies positive instances as negative [15].
Matthews correlation coefficient, which is a measure of the quality of binary (two-class) classification models. It takes into account TP, TN, FP and FN to provide a balanced assessment of the model’s performance, as shown in Eq. 3. The MCC ranges from \([-1,+1]\), where a value of \((+1)\) indicates a perfect classification, (0) indicates a random classification, and \((-1)\) indicates a completely wrong classification. MCC values closer to \((+1)\) indicate better performance of the classification model [3].
Micro-average is a method of aggregating the performance metrics across all classes in a multi-class classification problem. The formulas for micro-average precision (miP) (Eq. 4), recall (miR) (Eq. 5), and F1-score (miF1) (Eq. 6) are as follows [19]:

Macro-average, on the other hand, calculates the performance metrics for each class individually and then takes the average across all classes. The formulas for macro-average precision (maP) (Eq. 7), recall (maR) (Eq. 8), and F1-score (maF1) (Eq. 9) are as follows [19]:

where precision\(_i\), recall\(_i\), and F1-score\(_i\) represent the precision, recall, and F1-score of class i, and n is the total number of classes. By using micro and macro averaging, we can gain insights into the overall performance of the classification model, considering both the individual class performance and the overall performance across all classes.
4 Results and Discussion
In this section, we present a comprehensive analysis of various aspects related to convolutional neural networks (CNNs) and their fusion configurations. Firstly, we discuss the analysis for individual CNNs. Next, we delve into the evaluation of fusion configurations and CNN performance metrics. Furthermore, we investigate the fusion with optimal training CNN models. Lastly, we conduct a comprehensive analysis of resource consumption for the CNN models. By examining these four aspects, we gain a comprehensive understanding of the individual and fused CNN models, their performance metrics, optimal training configurations, and resource requirements. This knowledge enables us to make informed decisions and design more effective and resource-efficient CNN-based systems.
4.1 Analysis for Individual Convolutional Neural Networks
In this section, we evaluate the individual performance of the CNN models and their effectiveness in tackling the given task. This analysis provides insights into the strengths and weaknesses of each individual model. Table 2 present differents CNNs and configurations, such as learning rate (LR), in addition, performance metrics, including Matthews correlation coefficient (MCC), precision, recall, and F1-score. Each row represents a different CNN model, denoted by \(M_{1}\) to \(M_{7}\). Analyzing the results, it can be observed that different CNN models achieve varying levels of performance across the evaluated metrics. Among the models, \(M_{2}\) (DenseNet161) stands out with the highest precision, recall, F1-score, and MCC values. On the other hand, \(M_{5}\) (MobileNetV2) and \(M_{7}\) (VGG16) exhibit slightly lower performance in terms of precision, recall, F1-score, and MCC.
4.2 Analysis of Fusion Configurations and CNN Performance Metrics
In this section, we explore different fusion techniques and assess their impact on the overall performance of the system. Table 3 presents the results obtained from the fusion of multiple convolutional neural networks (CNNs), ensembles, using the average method. Each row in the table represents a different fusion scenario, denoted by \(F_{i}\), where multiple CNN models (\(M_{1}\) to \(M_{7}\)) are combined, where (Y) indicates the presence of a model in the fusion configuration, while (N) denotes the absence of that model. The table also includes the LR used for each fusion scenario. The evaluation metrics used to assess the performance of the fusion approach are precision, recall, F1-score, and MCC. Analyzing the results, it can be seen that different ensemble architectures and their fusion responses generate varying levels of performance in the evaluated metrics.
Among the merger scenarios, \(F_3\) displays the highest F1-score value for macro-average. This scenario combines specific CNN models (\(M_{2}\), \(M_{4}\), \(M_{6}\) and \(M_{7}\)) and achieves remarkable performance in correctly classifying positive and negative instances.
These scenarios show the effectiveness of ensemble methods to improve classification accuracy. Fusion \(F_1\) stands out, achieving relatively high values of accuracy, recall, F1-score and MCC, for micro average. This suggests that the combination of models \(M_2\), \(M_4\), \(M_6\) and \(M_7\) with an LR of \(\eta = 0.003\) leads to successful predictions with high accuracy and completeness. Considering the CNN models that appeared more frequently in the fusion experiments, the models \(M_2\), \(M_6\) and \(M_7\) were used in a greater number of experiments. This suggests that these models have a greater impact on the overall performance of the ensemble architectures. Table 4 shows the results of the CNN merger using a voting mechanism where each model in the ensemble makes an independent prediction, and the final prediction is based on the highest number of votes.
Upon analyzing the results, it is evident that the performance of the fusion models varies depending on the specific combination of CNN models used. \(F_2\) stands out as it achieves the highest values in terms of F1-score for macro (0.61582), F1-score for micro average (0.90892) and MCC (0.90115). This combination includes models \(M_1\), \(M_2\), \(M_3\), \(M_4\), and \(M_7\), indicating that these models contribute significantly to the overall performance of the fusion model.
4.3 Fusion with Optimal Training CNN Models
In this section, we explore the integration of specific CNN models to further enhance the system’s performance and achieve superior results. Table 5 presents the results of the fusion of trained CNN models using different combinations. In our experiments, in \(F_4^b\), considering the amount of CNNs that make up the fusion, we had the best performance compared to other combinations, considering both micro-average, F1-score (0.913) and MCC (0.905). Overall, the analysis of the fusion results indicates that the combinations involving \(M_2^b\) (DenseNet161), \(M_4^b\) (EfficientNet_b0), \(M_6^b\) (ResNet152), and \(M_7^b\) (VGG16) generally lead to better performance, with higher F1-scores and MCC values. The presence of \(M_1^b\) and \(M_5^b\) does not contribute significantly to the overall performance improvement.
4.4 Resource Consumption Analysis for CNN Models
In this section, we present Table 6, which provides details such as the CNN model name, GPU model used for execution, GPU RAM capacity, execution time in minutes, and the number of parameters for each model. To measure the execution time and GPU consumption, the timeit module and the psutil library were used, respectively. All network models, \(M_1\) to \(M_7\), utilize the Tesla V100-SXM2-16 GB GPU model. Additionally, the \(F_1^{b*}\) model [2] employs the Tesla V100-SMX2-16 GB, while fusion models \(F_2^b\) to \(F_7^b\), for Table 5 and Table 6, utilize the Nvidia A100-SXM-40 GB GPU model.
These data allow us to analyze the computational cost associated with achieving the results presented in Table 3, Table 4 and Table 5.
In Table 3, the \(F_{1}\) result, including \(M_{2}\), \(M_{4}\), \(M_{6}\), and \(M_{7}\), achieved the highest F1-score (0.91014) with the least number of models used for micro-average. The models were executed individually on the GPU, resulting in a total GPU consumption equal to the highest consumption among the individual models, which is 10 GB for \(M_{2}\). Thus, the proposed ensemble \(F_{1}\) with CNN model averaging has a maximum GPU consumption of 10 GB.
In Table 4, using the technique of fusion by vote, the ensemble \(F_{2}\) achieved the highest F1-score of 0.90892 for micro and F1-score of 0.61582 for macro. The set \(F_{2}\) consisted of models \(M_{1}\), \(M_{2}\), \(M_{3}\), \(M_{4}\) and \(M_{7}\). The GPU consumption for the set \(F_{1}\) corresponds to that of the model \(M_{2}\), which is 10 GB.
In Fusion with optimal training CNN models, as shown in Table 5, for \(F_1^{b*}\), the F1-score is 0.910, which matches our proposal in Table 3. The approach in [2], \(F_1^{b*}\), requires 15.8 GB of GPU, as the best models \(M_{2}^b\) and \(M_{6}^b\) are trained together. Building upon the combination of models proposed in [2], we introduce ensembles \(F_{2}^b\) to \(F_{7}^b\), with \(F_{4}^b\) achieving the best result. Figure 6 depicts a bubble chart illustrating that we have achieved comparable results (indicated by the blue and red bubbles) when compared to the fusion model \(F_1^{b*}\) in Table 5 (green bubble), as presented in [2]. Our proposal \(F_{1}\) in Table 3 attained the same results while utilizing 10 GB of GPU, which is \(36.7\%\) less than the consumption of [2] with 15.8 GB GPU. The purple bubbles demonstrate that our ensemble architectures using Fusion with optimal training CNN models obtain better results than [2], albeit at a higher GPU cost.
5 Conclusions
In conclusion, to aid in the diagnosis of diseases of the human gastrointestinal tract, through image classification, our objective was to propose ensemble architectures and efficient fusion techniques for CNNs, aiming to obtain better results than in the literature and to optimize computational resources.
A literature review and extensive experiments were conducted to validate the effectiveness of the proposed approach. The findings of this study highlight the significant contribution of DenseNet161 and ResNet152 to the fusion process in all experiments. Furthermore, our findings demonstrate a similar level of performance compared to the previous model, as indicated by an F1-score of 0.910 and MCC of 0.902. Remarkably, our approach achieves this performance using just 10 GB of GPU RAM, in contrast to the previous model’s requirement of 15.8 GB.
We were able to achieve a minimally higher F1 value of 0.913 and MCC of 0.905, but using 24.7 GB of GPU RAM. These findings contribute to our understanding of individual model performance, fusion techniques, and resource utilization, paving the way for the design and implementation of more efficient and effective deep learning-based systems in endoscopic scoring.
The main challenge was accessing sufficient computational resources. Future research should explore model compression techniques to reduce computational requirements. Additionally, further experiments and analysis are needed to identify optimal model combinations and refine the fusion process. Evaluating fusion alternatives on different datasets is crucial for assessing performance and generalizability.
Notes
- 1.
Available at: https://datasets.simula.no/hyper-kvasir.
- 2.
Available at: https://www.connectedpapers.com/main.
References
Ali, S., et al.: An objective comparison of detection and segmentation algorithms for artefacts in clinical endoscopy. Sci. Rep. 10(1), 2748 (2020)
Borgli, H., et al.: Hyperkvasir, a comprehensive multi-class image and video dataset for gastrointestinal endoscopy. Sci. Data 7(1), 283 (2020)
Chicco, D., Jurman, G.: The advantages of the Matthews correlation coefficient (MCC) over f1 score and accuracy in binary classification evaluation. BMC Genom. 21, 1–13 (2020)
Fukushima, K.: Neocognitron: a self-organizing neural network model for a mechanism of pattern recognition unaffected by shift in position. Biol. Cybern. 36(4), 193–202 (1980)
Hicks, S.A., Jha, D., Thambawita, V., Halvorsen, P., Hammer, H.L., Riegler, M.A.: The EndoTect 2020 challenge: evaluation and comparison of classification, segmentation and inference time for endoscopy. In: Del Bimbo, A., et al. (eds.) ICPR 2021. LNCS, vol. 12668, pp. 263–274. Springer, Cham (2021). https://doi.org/10.1007/978-3-030-68793-9_18
Iqbal, I., Walayat, K., Kakar, M.U., Ma, J.: Automated identification of human gastrointestinal tract abnormalities based on deep convolutional neural network with endoscopic images. Intell. Syst. Appl. 16, 200149 (2022)
Jha, D., et al.: Real-time polyp detection, localization and segmentation in colonoscopy using deep learning. IEEE Access 9, 40496–40510 (2021)
Jha, D., et al.: Medico multimedia task at mediaeval 2020: automatic polyp segmentation. arXiv preprint arXiv:2012.15244 (2020)
Jha, D., et al.: A comprehensive study on colorectal polyp segmentation with resunet++, conditional random field and test-time augmentation. IEEE J. Biomed. Health Inform. 25(6), 2029–2040 (2021)
Jha, D., et al.: Kvasir-SEG: a segmented polyp dataset. In: Ro, Y.M., et al. (eds.) MMM 2020. LNCS, vol. 11962, pp. 451–462. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-37734-2_37
Krizhevsky, A., Sutskever, I., Hinton, G.E.: Imagenet classification with deep convolutional neural networks. In: Advances in Neural Information Processing Systems, vol. 25, pp. 1097–1105 (2012)
LeCun, Y., Bengio, Y., Hinton, G.: Deep learning. Nature 521(7553), 436–444 (2015)
LeCun, Y., et al.: Backpropagation applied to handwritten zip code recognition. Neural Comput. 1(4), 541–551 (1989)
Naess, E., Thambawita, V., Hicks, S.A., Riegler, M.A., Halvorsen, P.: Pyramidal segmentation of medical images using adversarial training. In: Proceedings of the 2021 Workshop on Intelligent Cross-Data Analysis and Retrieval, pp. 33–38 (2021)
Sarkar, D., Bali, R., Sharma, T.: Practical Machine Learning with Python (2018). https://doi.org/10.1007/978-1-4842-3207-1
Sze, V., Chen, Y.H., Yang, T.J., Emer, J.S.: Efficient processing of deep neural networks: a tutorial and survey. Proc. IEEE 105(12), 2295–2329 (2017)
Sze, V., Chen, Y.H., Yang, T.J., Emer, J.S.: Efficient processing of deep neural networks. Synth. Lect. Comput. Archit. 15(2), 1–341 (2020)
Szegedy, C., Ioffe, S., Vanhoucke, V., Alemi, A.A.: Inception-v4, inception-resnet and the impact of residual connections on learning. In: Thirty-First AAAI Conference on Artificial Intelligence (2017)
Takahashi, K., Yamamoto, K., Kuchiba, A., Koyama, T.: Confidence interval for micro-averaged f 1 and macro-averaged f 1 scores. Appl. Intell. 52(5), 4961–4972 (2022)
Thambawita, V., et al.: An extensive study on cross-dataset bias and evaluation metrics interpretation for machine learning applied to gastrointestinal tract abnormality classification. ACM Trans. Comput. Healthc. 1(3), 1–29 (2020)
Acknowledgements
This study was financed in part by Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001* and Conselho Nacional de Desenvolvimento Científico e Tecnológico (grant 306436/2022-1). In addition, it had the support of the Instituto Federal do Triângulo Mineiro e Universidade Federal de Uberlândia.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Costa, C.L., Lima, D.A., Zorzo Barcelos, C.A., Travençolo, B.A.N. (2023). Ensemble Architectures and Efficient Fusion Techniques for Convolutional Neural Networks: An Analysis on Resource Optimization Strategies. In: Naldi, M.C., Bianchi, R.A.C. (eds) Intelligent Systems. BRACIS 2023. Lecture Notes in Computer Science(), vol 14196. Springer, Cham. https://doi.org/10.1007/978-3-031-45389-2_8
Download citation
DOI: https://doi.org/10.1007/978-3-031-45389-2_8
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-45388-5
Online ISBN: 978-3-031-45389-2
eBook Packages: Computer ScienceComputer Science (R0)