Abstract
Advances in textual classification can foster quality in existing clinical systems. Our research explored experimentally text classification methods applied in non-synthetic oncology clinical notes corpora. The experiments were performed in a dataset with 3,308 medical notes. Experiments evaluated the following machine learning and deep learning classification methods: Multilayer Perceptron Neural network, Logistic Regression, Decision Tree classifier, Random Forest classifier, K-nearest neighbors classifier, and Long-Short Term Memory. An experiment evaluated the influence of the corpora preprocessing step on the results, allowing us to identify that the classifier’s mean accuracy was leveraged from 26.1% to 86.7% with the per-clinical-event corpus and 93.9% with the per-patient corpus. The best-performing classifier was the Multilayer Perceptron, which achieved 93.90% accuracy, a Macro F1 score of 93.61%, and a Weighted F1 score of 93.99%.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Today’s world is moving toward changing healthcare from reactive and hospital-centered to preventive and personalized treatment approaches [1]. Therefore, physicians must be updated with these new treatments and patients’ health information. Furthermore, decision-making is affected by a broad range of parameters. For instance, different information sources are used to define oncology diagnosis and treatment options [2]. In this context, the information accumulated in a health institution’s database can prove valuable to help physicians decide on the appropriate course of action in specific cases [3]. Nevertheless, it is not feasible for health professionals to manually evaluate and analyze these resources due to the enormous effort it will demand, considering the size of these databases [4], and the exponential growth in medical information [5].
Due to the rapid growth of electronic information, most health information about patients is found as unstructured data, i.e., narrative text or free text [3]. Therefore, the unstructured format of the enormous amount of medical data is the primary motive of recent research initiatives aiming to extract information from medical notes [6,7,8,9].
Considering these challenges, tools are needed to support healthcare professionals in their activities beyond a healthcare system’s restructuring. Clinical Decision Support Systems (CDSS) provide additional assistance, synthesizing and integrating patient-specific information, performing complex evaluations, and presenting the results to clinicians in an adequate time [10]. In addition, CDSS offers assistance to overcome the difficulties in dealing with massive amounts of information [11,12,13].
This work focuses on the text classification task, covering the steps to create and process a clinical corpus for the experiments and comparing evaluations with machine learning and deep learning approaches. An oncology clinical notes corpus was created, preprocessed, and transformed to be used by machine learning and deep learning methods. This corpus was obtained from a real-world oncology clinic, de-identified to preserve the patient and professional’s identification, and is entirely composed of Brazilian Portuguese language texts. In addition, this work implemented several machine learning and deep learning methods for text classification, compared their performance, and evaluated their results.
Related works about text classification and information extraction methods in oncology clinical notes were studied to understand how to better deal with unstructured data. The papers used one or a combination of natural language processing (NLP), machine learning, and deep learning methods. The related works study indicates text classification and information extraction as essential tasks to deal with unstructured data in healthcare. Several related works used domain-specific corpora. However, none used a corpus with non-synthetic medical notes from the oncology healthcare-specific domain in the Brazilian Portuguese language. Furthermore, most papers used machine learning or deep learning methods to consider the text classification task. Just a few of them presented a comparison between several machine learning and deep learning methods.
Therefore, the main objective of this work consists of applying text classification approaches to support healthcare professionals regarding diagnosis decisions. The survey provided insight into crucial aspects of the area, such as that patients’ clinical data are often inputted in a free-text format. There was a lack of structured data repositories on this topic. This topic was described as an essential problem in most studied papers considering that CDSS needs data in a structured format. The corpora creation is an essential element in this research and constitutes one of the contributions of this work because it was created from non-synthetic data with specific preprocessing needs. Other contributions presented in this work are the corpus de-identification and enrichment process, described in the following sections. The evaluation and performance comparison of several machine learning and deep learning classifiers also brings contributions from this work.
2 Material and Methods
This section presents the methodological aspects adopted in this work. This work aims to evaluate text classification techniques to support health professionals’ needs regarding diagnosis decisions. Therefore, text classification experiments were conducted using machine learning and deep learning approaches. These techniques were applied in Brazilian Portuguese clinical notes corpora obtained from an EHR system in the oncology domain.
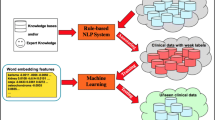
Figure 1 shows the general view containing the elements of the proposed approach for this work. This context is derived from real-world observation, considering actual medical clinics, and expresses health professional needs.
The overall process starts with the creation of a record of a medical note by the healthcare professional. This situation generates textual medical records and structured clinical information in the real-world observed cases. In this step, the healthcare professionals use the Oncology EHR system [14] to input their observations about the patient, which are recorded in the system’s database. These observations can be composed of free-text and structured data, and both data types are used together to achieve better results.
The upper part of Fig. 1 describes the primary steps for exploring possible answers to the research question in this work. First, developing a flow of operations that starts with the health professional’s assistance was evaluated, as identified in the flow “Load new medical textual records” used together with a support service, applying the classification models studied. The application of the models will generate support for determining a response with suggestions for framing and similar contexts, as represented by the “Diagnosis and treatment recommendation” flow that the physician will use. The generation of new information about each patient’s clinical event is stored in the Oncology EHR system’s database, as indicated in the “Record medical notes” flow.
This initial flow consists of a vision of the future use of the system by physicians and healthcare professionals. The other items in Fig. 1 were studied and experimented.
The first necessary step to generate the corpora and to use the proposed approach in the future was to anonymize and export these records. First, a de-identification process was applied to anonymize the data, preventing the personal identification of each patient and professional. After de-identification, the records were exported, creating the corpus used in text classification approaches. In this step, the textual data was exported along with some structured clinical information.
The corpus was annotated, taking advantage of free-text data labels generated by the medical staff when using the EHR system and stored in the database. Therefore, this corpus could be considered for some of the classification approaches tasks, such as the training tasks.
After the corpus creation and annotation, it was enriched with some structured clinical information. As mentioned previously, the EHR system stores the data as a free-text type or structured type. Some of these structured data are the diagnosis and treatment information used to enrich the corpus.
The final step involves all the preprocessing steps necessary to train the evaluated classification algorithms. Before training the artificial neural network model, the corpus was preprocessed as described in Sect. 2. In this step, the corpus annotations and enriched data were assessed. When necessary, textual features were extracted. After that, the corpus and its features were used as input to train the following Artificial Neural Network (ANN) models: Multilayer Perceptron (MLP) and Long short-term memory (LSTM). Part of the corpus was reserved for testing to evaluate each ANN model’s performance.
We developed experiments combining approaches such as corpus format, new corpus with updated information, document or paragraph processing levels, and different machine learning and deep learning classifiers (as described in Sect. 3). The experiments’ results are evaluated in Sect. 3.3.
As a result of the work overviewed above, creating a corpus with non-synthetic oncological medical notes and implementing a de-identification and enrichment process of the corpus are highlighted. In addition, the evaluation and performance comparison of a machine learning and a deep learning classifier are also highlighted as contributions of this work.
In the following sections, each component or step involved in the general approach of Fig. 1 is detailed.
The Machine Learning and Deep Learning architectures will be described, along with some parametrization aspects, complementary information on the feature engineering, and the general view of the planned experiments set.
The additional corpora preprocessing, the feature extraction details, and the model specifications are described in the following subsections.
Corpus Preprocessing. After the corpora creation process, it was necessary to preprocess them to normalize the text. The same preprocessing was performed for both per-clinical-event and per-patient corpora types. The text analysis and preprocessing techniques applied are described below.
Before the text preprocessing, it was possible to observe that a small group of diagnoses concentrates on the most frequent occurrences. Hence the diagnoses with less than 50 occurrences were joined into a single group called “Outros” (“Others”). Furthermore, to evaluate the neural network’s performance according to the dataset sparsity, a new version of the dataset was created with the 12 most occurring diagnoses.
The following tasks were performed:
-
Tokenization: split the text into tokens that correspond to words;
-
Stop-words filtering: removal of most common words in the Brazilian Portuguese language, punctuation, and special characters;
-
Case folding: conversion all words to lowercase.
An additional manual analysis was done on the corpus in the per-clinical-events corpora. The text was assessed to understand how it could be transformed to improve ANN algorithms. Repeated medical notes were removed from several annotation labels. These notes could weaken the representation strength of the corpus.
A complementary experiment was performed to evaluate how this step leveraged the results of the classifiers. As described in Sect. 3.1, a significant improvement was achieved by applying this step.
Feature Extraction. The text from medical notes must be transformed into a structure that the classifiers could use. For that reason, the Bag-of-Words (BoW) method was used, which is a representation that turns arbitrary text into fixed-length vectors by counting how many times each word appears. This representation is useful to be used by the classifier algorithms.
This work used the medical notes bag-of-words (BoW) to extract the features to be used in machine learning and deep learning training. In the per-clinical-event corpora, the BoW was generated for each medical note. Likewise, the per-patient corpora were generated for each patient with all their medical notes.
Before creating the medical notes BoW, the text was normalized as described in the preprocessing Sect. 2. This preprocessing step aims to reduce the number of useless words, special characters, and punctuations, which would not make a difference in the classifier model’s training. It also helped to reduce the computational effort to create the BoW.
The BoW applied to the medical notes resulted in a sparse representation, i.e., the vector sequence of numbers representing each word contained too many zeros. The Principal Component Analysis (PCA) technique was applied to reduce the data sparsity. The PCA technique converts a set of observations of possibly correlated features into values of linearly uncorrelated features. The PCA with 500 features was used.
Machine Learning and Deep Learning Architectures. This work applied machine learning and deep learning methods for the text classification task, comparing their results. Several machine learning classification algorithms were applied to evaluate which one had the best performance. Furthermore, an LSTM deep learning algorithm was also applied to compare traditional machine learning and a deep learning recurrent neural network.
The following Machine Learning algorithms were evaluated: Multilayer Perceptron (MLP) neural network [15]; Logistic Regression [16]; Decision Tree classifier [17]; Random Forest classifier [18]; K-Nearest Neighbors (KNN) classifier [19]. Furthermore, a Long-short Term Memory (LSTM) deep learning experiment was also performed. The machine learning classification and the deep learning architecture used are described, regarding their algorithms and their theoretical background, in the following papers [20,21,22].
The datasets were divided into two groups: one with 80% of the data to train the models and 20% of the data to test the models. The data were shuffled to keep the categories’ proportion, and then they were divided as aforementioned.
The machine learning algorithms were implemented using scikit-learnFootnote 1, and the deep learning LSTM was performed using Keras libraryFootnote 2, both were implemented in Python. In the first set of experiments, seven tests were performed, with the following architecture details: an MLP with one hidden layer with 500 neurons; an MLP with two hidden layers with 800 and 500 neurons; a Logistic Regression classifier; a Decision Tree with a maximum of twenty levels and three samples by leaf; a Random Forest with a maximum of twenty levels and three samples by leaf; an Extra Trees with a maximum of twenty levels and three samples by leaf; a KNN classifier with a unitary K.
The second set of experiments was performed this time with the 12 most occurring diagnoses in the dataset. For this experiment, the best-performing machine learning was selected to compare with an LSTM deep learning recurrent neural network. The machine learning algorithm had the following architecture: an MLP with two hidden layers with 800 and 500 neurons. The deep learning algorithm had the following architecture: an LSTM with the library Keras built on top of Tensorflow in a python implementation, in which the parametrization used was composed of Batch size 128, the Dropout rate of 0.2, validation split of 0.2, Optimizer with adam, Loss measure was categorical cross-entropy. Also, to prevent overfitting, EarlyStopping was used. Standard values for the parameters were used in these experiments, as described in the literature. All the described models were evaluated using standard metrics indicated in the literature, such as accuracy and macro and weighted F1 scores.
3 Experiments and Results
Several machine learning classifiers and a deep learning recurrent neural network were applied in this work’s experiments. The main objective of these experiments was to address the main research question and identify a possible workflow to use the dataset and text classification algorithms to evaluate potential support for healthcare professionals.
The dataset used was composed of an arrangement of the available options, using the per-clinical-event and the per-patient versions. The complete dataset with several machine learning text classification algorithms was used in the first step. Both datasets (per clinical event and per-patient) were used in this experiment. In the second step, a new experiment was carried out involving the per-patient dataset and the algorithms MLP and LSTM. The per-patient dataset was chosen in the second step because all patient’s clinical notes were joined into a single record, which would perform better considering the LSTM’s ability to process entire sequences of data. All methods were executed on a partition, then repartitioned and reran all methods to get the mean Accuracy.
Therefore, two main experiments were performed: a) Machine learning - several machine learning classifiers have been experimented with and their performance compared (described in Sect. 3.1); b) Deep learning - an experiment with a deep learning recurrent neural network was performed (described in Sect. 3.2).
To measure performance in the experiments, we use different metrics in this study: Accuracy, macro, and weighted F1 score [20]. The predicted output as True Positive (TP) indicates text classified as correct, True Negative (TN) when classified incorrectly. False Positive (FP) if a text correctly indicates not belonging to the class. Similarly, a False Negative (FN) is the text is classified incorrectly.
The Accuracy describes the overall performance of the classifier and mathematical Accuracy expressed as below in Eq. (1):
The F1-score It is a harmonic measure between precision and sensitivity, expressed in the Eq. (2):
The Macro F1-score is defined as the mean of class-wise/label-wise F1-scores, in Eq. (3), where i is the class/label index and N the number of classes/labels:
In weighted-average F1-score, we weight the F1-score of each class by the number of samples from that class, such as in Eq. (4), where i is the class index and N the number of classes and S the number of elements in the class.
3.1 Machine Learning Experiments
The per-clinical-event corpus (described in Sect. 2) of the smallest clinic in the preprocessed and raw versions were used to perform the classifiers. This is the clinic database and contains 3.308 clinical notes, and 397 distinct patients. The preprocessed dataset was used first with the machine learning classifiers, described in Sect. 2.
To perform the experiments, the dataset was randomly divided into two parts: 80% for training and 20% for testing. A shuffle method was used to generate these two parts, and a different set of training and testing datasets were created each time it was performed. Hence different classifiers’ metrics were obtained, but it always kept the performance order.
The mean accuracy, Macro F1 score, and Weighted F1 score of each classifier are presented in Table 1. These experiments were performed to evaluate which machine learning classifier had the best performance. According to Table 1, the MLP 2 classifier achieved the best accuracy, Macro F1, and Weighted F1 scores. These results are evaluated in Sect. 3.3.
An additional experiment evaluated how the dataset’s structure and preprocessing step (Sect. 2) leveraged the classifiers’ performance. This experiment used the same dataset from the Clinic with the smalest dataset. The preprocessed and raw versions of the per-clinical-event dataset, plus the preprocessed per-patient dataset, were used with the best performance classifier. According to Table 1, the MLP 2 classifier had the best performance and was used in this experiment.
Table 2 presents the mean accuracy of the MLP 2 classifier with preprocessed and raw versions of the per-clinical-event dataset, plus the preprocessed per-patient dataset.
These results are evaluated in Sect. 3.3, exploring the improvements obtained with the integration of the clinical events in a more complete view of the patient history. As it can be observed in Table 2, the mean accuracy improves with more complete patient data.
3.2 Deep Learning Experiments
In this set of experiments, the following machine learning and deep learning classifiers were tested:
-
The machine learning classifier that best performed (the MLP 2), according to Sect. 3.1;
-
An LSTM (Long-short term memory) deep learning recurrent neural network.
The per-patient corpus (described in Sect. 2) of the smallest clinic in the preprocessed version was used to perform the classifiers. The per-patient corpus was chosen because all patient’s clinical notes were joined into a single registry, which would perform better considering the LSTM ability to process entire sequences of data.
Table 3 presents the mean accuracy, Macro F1 score and Weighted F1 score of the MLP 2 and the LSTM classifiers. These results are evaluated in Sect. 3.3.
3.3 Results Evaluation
Several experiments were performed to understand the behavior of the selected machine learning and deep learning classifiers with the corpora created and preprocessed (Sect. 2).
First, a set of experiments with seven machine learning classifiers were performed with the per-clinical-event corpus, according to Sect. 3.1. Considering the mean accuracy, Macro F1, and Weighted F1 scores, the classifier that best performed was the MLP 2, as seen in Fig. 4.
Preparing the corpus to be used in machine learning and deep learning classifiers was an important step. An experiment was performed with the MLP 2 classifier, the preprocessed and raw versions of the per-clinical-event dataset, and the preprocessed per-patient dataset (as described in Sect. 2). The performance of each is presented in Table 2. This table shows a significant improvement in the MLP 2 classifier performance with the preprocessing of the dataset (Sect. 2), in both per-clinical-event and per-patient datasets. Furthermore, the per-patient corpus performed better than the per-clinical-events corpus. For that reason, the next experiments used the per-patient corpus.
After the evaluation of the machine learning classifier that best performed, the MLP 2 classifier was selected. A new experiment was performed, to compare the MLP 2 classifier with an LSTM recurrence neural network, using the preprocessed per-patient corpus (as described in Sect. 3.2). Figure 5 shows that the MLP 2 performed better than the LSTM classifier, even though the latter is a more recent neural network. This result can be associated with the fact that this experiment used the smallest per-patient corpus, and deep learning algorithms perform better on large datasets.
4 Conclusion
With a large amount of information generated daily in healthcare, it is unsuitable for humans to process it manually. Furthermore, much of this information is recorded as unstructured data, transforming it into a hard task. It is necessary to develop tools to help healthcare professionals to deal with it, automate the classification and extraction of information from medical notes, and enable this information to be processed by machines.
This work created the corpus based on non-synthetic oncological medical notes from an Oncology EHR system database. This corpus creation process allowed the identification of various preprocessing steps (as described in Sect. 2), and specific term treatment to improve overall results. The experiments achieved good accuracy, especially MLP machine learning and LSTM deep learning methods, showing possibilities of using these resources for medical notes text classification. Therefore, this set of tasks provided strong support indicating resources and processes to text classification in the specific context of health professional support.
Artificial Intelligence, special machine learning, and deep learning algorithms have been widely applied in several industries, sometimes surpassing human accuracy. In the healthcare industry, several processes can be improved by AI, leveraging healthcare professionals. In the oncology area, diagnosis and treatment decision-making is one of these complex processes that AI algorithms can aid.
Considering this work’s development, the following list of future steps is suggested:
-
Create larger corpus with the medical notes from several oncology clinic databases and conduct new experiments;
-
Enhance the corpus preprocessing step by removing low-frequency words, spell checker, replacing acronyms and abbreviations by its standard word;
-
Create a domain-specific word embedding from the corpora and apply it with the classifiers. Another option is to use general-purpose word embeddings and fine-tune it with the corpora;
-
Improve the enrichment process with more structured data available from the Oncology EHR system, such as prescribed medications, patient’s problems, and allergies;
-
Tune the implemented classifiers and try different versions, such as a Bidirectional LSTM (Bi-LSTM);
-
Integrate the implemented classifiers with the Oncology EHR system to obtain feedback from the healthcare professionals about their accuracy, and suggest the diagnosis based on the patient’s clinical history.
Notes
References
Sabra, S., Alobaidi, M., Malik, K.M., Sabeeh, V.: Performance evaluation for semantic-based risk factors extraction from clinical narratives. In: IEEE 8th Annual Computing and Communication Workshop and Conference (CCWC). Las Vegas, NV, USA, vol. 2018, pp. 695–701 (2018). https://doi.org/10.1109/CCWC.2018.8301742
Glatzer, M., Panje, C.M., Sirén, C., Cihoric, N., Putora, P.M.: Decision making criteria in oncology. Oncology 98(6), 370–378 (2020). Epub 2018 Sep 18. PMID: 30227426. https://doi.org/10.1159/000492272
Reyes-Ortiz, J.A., González-Beltrán, B.A., Gallardo-López, L.: Clinical decision support systems: a survey of NLP-based approaches from unstructured data. In: 26th International Workshop on Database and Expert Systems Applications (DEXA). Valencia, Spain vol. 2015, pp. 163–167 (2015). https://doi.org/10.1109/DEXA.2015.47
Alemzadeh, H., Devarakonda, M.: An NLP-based cognitive system for disease status identification in electronic health records. In: 2017 IEEE EMBS International Conference on Biomedical & Health Informatics (BHI), Orlando, FL, USA, pp. 89–92 (2017). https://doi.org/10.1109/BHI.2017.7897212
Meskó, B.: The guide to the future of medicine: technology and the human touch. In: Webicina KFT (2014)
Zhang, R., Ma, S., Shanahan, L., Munroe, J., Horn, S., Speedie, S.: Automatic methods to extract New York heart association classification from clinical notes. In: 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Kansas City, MO, USA, pp. 1296–1299 (2017). https://doi.org/10.1109/BIBM.2017.8217848
Chen, X., Xie, H., Wang, F., et al.: A bibliometric analysis of natural language processing in medical research. BMC Med. Inform. Decis. Mak. 18(Suppl 1), 14 (2018). https://doi.org/10.1186/s12911-018-0594-x
Shickel, B., et al.: Deep EHR: a survey of recent advances in deep learning techniques for electronic health record (EHR) analysis. IEEE J. Biomed. Health Inform. 22(5), 1589–1604 (2017)
Kreimeyer, K., et al.: Natural language processing systems for capturing and standardizing unstructured clinical information: a systematic review. J. Biomed. Inform. 73, 14–29 (2017). Epub 2017 Jul 17. PMID: 28729030; PMCID: PMC6864736. https://doi.org/10.1016/j.jbi.2017.07.012
Hunt, D.L., Haynes, R.B., Hanna, S.E., Smith, K.: Effects of computer-based clinical decision support systems on physician performance and patient outcomes: a systematic review. JAMA 280(15), 1339–1346 (1998). PMID: 9794315. https://doi.org/10.1001/jama.280.15.1339
Bucur, A., van Leeuwen, J., Cirstea, T.C., Graf, N.: Clinical decision support framework for validation of multiscale models and personalization of treatment in oncology. In: 13th IEEE International Conference on BioInformatics and BioEngineering, Chania, Greece, pp. 1–4 (2013). https://doi.org/10.1109/BIBE.2013.6701695
Polpinij, J.: The cancerology ontology: designed to support the search of evidence-based oncology from biomedical literatures. In: 24th International Symposium on Computer-Based Medical Systems (CBMS). Bristol, UK, pp. 1–6 (2011). https://doi.org/10.1109/CBMS.2011.5999168
Wang, Y., et al.: Clinical information extraction applications: a literature review. J. Biomed. Inform. 77, 34–49 (2018). ISSN 1532–0464. https://doi.org/10.1016/j.jbi.2017.11.011
InterProcess: InterProcess Gemed Oncology - Oncological management system (2019). www.interprocess.com.br/en/gemed-oncology/
Naraei, P., Abhari, A., Sadeghian, A.: Application of multilayer perceptron neural networks and support vector machines in classification of healthcare data. In: Future Technologies Conference (FTC). San Francisco, CA, USA, vol. 2016, pp. 848–852 (2016). https://doi.org/10.1109/FTC.2016.7821702
Lemon, S.C., Roy, J., Clark, M.A., Friedmann, P.D., Rakowski, W.: Classification and regression tree analysis in public health: methodological review and comparison with logistic regression. Ann. Behav. Med. 26(3), 172–181 (2003). PMID: 14644693. https://doi.org/10.1207/S15324796ABM2603_02
Lavanya, D., Rani, K.U.: Ensemble decision tree classifier for breast cancer data. Int. J. Inf. Technol. Convergence Serv., 2(1), 17 (2012)
DuBrava, S., et al.: Using random forest models to identify correlates of a diabetic peripheral neuropathy diagnosis from electronic health record data. Pain Med. 18(1), 107–115 (2017). PMID: 27252307. https://doi.org/10.1093/pm/pnw096
Tayeb, S., et al.: Toward predicting medical conditions using K-nearest neighbors. In: 2017 IEEE International Conference on Big Data (Big Data), Boston, MA, USA, pp. 3897–3903 (2017). https://doi.org/10.1109/BigData.2017.8258395
ul Haq, A., Li, J.P., Memon, M.H., Nazir, S., Sun, R.: A hybrid intelligent system framework for the prediction of heart disease using machine learning algorithms. Hindawi Mobile Inf. Syst. (2018). https://doi.org/10.1155/2018/3860146
Haq, A.U., et al.: Intelligent machine learning approach for effective recognition of diabetes in E-Healthcare using clinical data. Sensors 20, 2649 (2020). https://doi.org/10.3390/s20092649
Tai, K.S., Socher, R., Manning, C.D.: Language processing, improved semantic representations from tree structured long short-term memory networks. In: Proceedings of the 53rd Annual Meeting of the Association for Computational Linguistics and the 7th International Joint Conference on Natural Language Processing of the Asian Federation of Natural, pp. 1556–1566 (2015)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
da Silva, D.P., Fröhlich, W.d.R., Schwertner, M.A., Rigo, S.J. (2023). Clinical Oncology Textual Notes Analysis Using Machine Learning and Deep Learning. In: Naldi, M.C., Bianchi, R.A.C. (eds) Intelligent Systems. BRACIS 2023. Lecture Notes in Computer Science(), vol 14196. Springer, Cham. https://doi.org/10.1007/978-3-031-45389-2_10
Download citation
DOI: https://doi.org/10.1007/978-3-031-45389-2_10
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-45388-5
Online ISBN: 978-3-031-45389-2
eBook Packages: Computer ScienceComputer Science (R0)