Abstract
Constructive Machine Learning (CML) is a research field that uses algorithms to generate new instances, similar but not identical to existing ones. It has been widely used to assist the discovery of new drug-like molecules. This is very challenging, given that the search space is discrete, unstructured and enormous. In this work we use CML to learn the intrinsic rules of datasets of molecules to generate novel ones. The chosen CML methods can be divided in two sub groups, text-based and graph oriented. Considering different possibilities to evaluate the methods and the generated molecules, we propose classifying generated molecules in a taxonomy, using a hierarchical multi-label classifier previously trained in a dataset of molecules with known taxonomy information. In this way, it is possible to predict properties and verify the relevance of the generated molecules to existing taxonomies. We also propose a hierarchical diversity measure to compare groups of molecules based on their taxonomy information. The measure showed coherent results and is faster to calculate than the commonly used external diversity measures.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
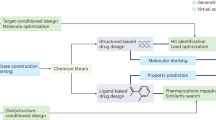
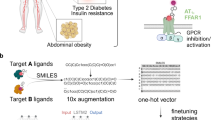
Similar content being viewed by others
1 Introduction
One of the objectives of Medicinal Chemistry is to discover new molecules with drug-like characteristics. This is challenging, since searching for molecules with desired chemical properties involves a huge and discrete space. The number of possible candidates is overwhelming and makes impossible to brute force through all possible combinations. About \(10^8\) substances have been synthesized, whereas the range of potential drug-like molecules is estimated to be between \(10^{23}\) and \(10^{60}\) [14]. Other problem is that the space is unstructured, i.e., adding, removing or changing atoms and bonds of a molecule changes its properties in an inconstant and hard to predict way. Despite improvements in the synthesis and test areas, those procedures are still time consuming and expensive. The discovery process can take 10–15 years with an average cost of US$2.5 billion [9], and one-third of the time and money are spent during the early stages of the process.
Machine Learning (ML) is relevant to manage the volume and complexity of chemical information. A single database (such as PubChem [18]) has more than 96 million chemical structures, with an even larger number of annotations (such as synonyms and properties), targets (drugs with which a molecule interacts), action (how the molecule interacts with its targets), and status within the regulatory-approval process [31]. ML seeks to generalize patterns observed in datasets and predicts unknown characteristics of unseen data. It has been applied in drug design to estimate properties of molecules and predict interactions with biological targets, considerably accelerating drug discovery [6].
Constructive Machine Learning (CML) generates instances similar, but no identical, to known instances. It has shown outstanding results in many areas and is gaining increasing attention [12]. In recent years, CML has been applied to generate new molecules from scratch based on existing ones [24]. Four challenges are observed when applying CML to de novo drug design [10]: i) definition of molecules representation; ii) architecture selection; iii) evaluation of approaches for molecules generation and optimization; and iv) design of a reward function, crucial for practical applications of reinforcement learning. Although there is a variety of proposals to overcome these challenges, there are still open problems.
Traditional ML classifiers were developed for binary or multi-class problems. They are not suitable for real-world problems where instances are classified into many classes simultaneously. An example is Hierarchical Multi-label Classification (HMC), where instances are classified into different classes in a taxonomy. These problems require hierarchical multi-label classifiers [30].
Many works validate molecules using classifiers to predict desired molecules properties. However, there are no works applying HMC to classify de novo molecules into known properties within a taxonomy. This is important, since we can directly identify general and specific properties, efficiently comparing groups of molecules. Thus, we propose to evaluate our molecules by classifying them into a taxonomy, using a hierarchical multi-label classifier previously trained using molecules with known taxonomy information. To generate molecules, we use the Generative Toolkit for Scientific Discovery (GT4SD) [28], and classify them using a decision tree [30] trained in the ChEBI molecules dataset [8]. For evaluation, we propose a hierarchical distance to compare groups of molecules based on their taxonomy information predicted by the hierarchical decision tree. Our main contributions are: i) the study and comparison of CML models for molecules generation considering their taxonomy; ii) the creation of a library to process the ChEBI dataset for hierarchical classification; iii) the proposal of a new measure to compare groups of molecules based on their taxonomy.
The remainder of this paper is organized as follows. Section 2 presents related works; Sect. 3 reviews fundamental related concepts; Results are presented and discussed in Sect. 4; Finally, Sect. 5 presents conclusions and future works.
2 Related Works
Recurrent Neural Networks (RNN) were applied to learn intrinsic molecules rules [5], comparing distributions of descriptors to verify if they were similar to existing ones. RNNs were also applied to generate molecules from scratch and from scaffolds [15]. Fine tuning was performed in smaller specific datasets with molecules with known biological targets. Principal Component Analysis was applied to the descriptors of the training data and the generated data, showing that the distributions were very close.
Reinforcement Learning (RL) has been extensively investigated due to its potential applications in drug discovery. In a recent study, the combination of RL and Quantitative Structure Activity Relationship (QSAR) was explored as a reward function for identifying active molecules against the Dopamine Receptor Type 2 (DRD2) [24]. The study used RNN and QSAR model to generate structures. The results showed that more than 95% of the structures generated were active against DRD2, demonstrating the potential of RL in drug discovery.
A method called ORGAN combines Generative Adversarial Networks (GANs) with high Quantitative Estimate of Drug-Likeness (QED) [26]. It has three parts: a generator, a discriminator, and QED with Lipinski’s rule-of-five as reward function. Variational Autoencoders (VAEs) were also investigated [14]: i) sampling molecules next to known ones; ii) interpolating points from known molecules and converting them back; iii) using a property predictor to search optimal regions, converting points with high values into molecules. The first approach chooses a molecule as reference and samples points around it. The second chooses two molecules as references, smoothly interpolating molecules similar to them. The third approach showed that VAEs trained with the property predictor led to regions where the chosen property is maximized. Adversarial Autoencoders (AAEs) [17] were compared with VAEs, showing that VAEs have a greater coverage. Allowing to trade-off between coverage of the original dataset with generated samples and reconstruction quality, AAEs obtained results similar to VAEs.
Graph Convolutional Policy Network (GCPN) [34] used a generation process guided towards specified desired objectives, restricting the output space based on underlying chemical rules. For goal-directed generation, graph representation, reinforcement learning and adversarial training were used, extended and combined into an unified framework.
A flow-based auto-regressive model (GraphAF) was proposed [27], combining Auto Regressing Flows [25] and graph representation learning. Motifs were also used [21], which are fragments of molecules inferred from data. VAEs were extended to work with graph molecular representation, and generated molecules including motifs, a bond or a atom per iteration. The proposal, called MoLeR (learning to extend molecular scaffolds with structural motif), was argued to be faster than baseline methods. The method uses an off-the-shelf molecular optimization, combining state-of-the-art methods in unconstrained optimization.
From the reviewed works, we generated molecules with the following methods: VAE [14], ORGAN [26], AAE [17], MoLeR [21], GCPN [34] and GraphAF [27].
3 Methodology
This section presents concepts, algorithms and data used in our experiments. Section 3.1 introduces the concepts of molecules properties and Sect. 3.2 presents three molecular representations used in our experiments. The GT4SD library and the Clus system, used to generate and classify molecules, are described in Sects. 3.3 and 3.4. Section 3.5 provides information about the ChEBI taxonomy, used to train our classifier and validate molecules. The next two sections present metrics used to compare datasets of molecules. Section 3.6 describes the internal and external diversity measures, while Sect. 3.7 explains our proposed hierarchical distance. Section 3.8 presents an overview of our methodology.
3.1 Molecules Properties
Each molecule has i) its own structure, i.e., how its atoms are organized; ii) its physical properties (descriptors), such as its weight and solubility; and iii) its biological properties, related to effects in biological targets [4]. Cheminformatics tries to predict molecules physical properties using their chemical structure, and to predict biological targets using known properties [31].
Although we can not say for sure if a molecule has pharmaceutic characteristics, we can observe physicochemical and structural properties within certain ranges. Over 90% of molecules that reach the phase II clinical status have four properties: i) mass lower than 500; ii) logP lower than 5; iii) number of Hydrogen bond donors lower than 5; and iv) number of Hydrogen bond receptors lower than 10. A molecule satisfying these restrictions is a strong candidate to be a drug. Although these are not the only properties for the molecule to be a drug, they have shown to be very influential, and are known as Rule of Five (RO5) (or Lipinski rule) [19]. Even though the RO5 is predictive of oral bioavailability, 16% of oral drugs violate at least one of the criteria and 6% fail on two or more.
An important estimated molecule property is the Quantitative Estimate of Drug-Likeness (QED), which is used to assess molecular target druggability by prioritizing bioactive compounds, directing the tests to most promising molecules [4]. Quantitative Structure Activity Relationship (QSAR) [23] indicates how a chemical compound interacts with biological targets. It can be used as objective function, helping to predict how promising a molecule is to be a drug.
In our experiments, we trained a hierarchical multi-label classifier using molecules extracted from the ChEBI database, having as features the above properties and other descriptors. These same features were extracted from the molecules we generated using CML. These molecules were then classified using the hierarchical classifier. Since molecules with similar functions are close in the feature space [31], we evaluated our generated molecules using similarity measures, comparing groups of molecules generated by different CML algorithms.
3.2 Molecules Representation
The representation of a molecule is a key factor to determine how CML models work and how similarities are evaluated. The Simplified Molecular Input Line Entry System (SMILES) is a two dimensional representation that uses ASCII symbols: letters for atoms, = and # for bonds, parenthesis for ramifications and numbers for cycles; simple bonds and hydrogen atoms are sub intended. A positive characteristic of SMILES is its ability to capture order and relation of atoms. However, it allows to represent a same molecule in many different ways, which difficulties comparisons. A same molecule can have different similarity scores depending on the chosen atom to start its sequence. Also, a single character perturbation in the representation can lead to changes in the underlying molecular structure or even invalidate it [32]. We used this representation for some CML models in the generation stages, and to store molecules.
Fingerprints uses an array of predefined size, where each position indicates the presence or the absence of a molecular structure. Its positive characteristics are that there is only one possible encoding for each molecule, and it is simple to compare vectors with fixed sizes. However, this representation cannot capture the order neither the relationship between the encoded structures. Therefore, an encoding can be related to numerous molecules, making it impossible to reverse the codification [7]. We used this representation to compare molecules.
The graph representation uses vertices and edges to represent atoms and bonds, providing high coverage of the chemical space, and conveying various chemical features. It is argued that it is the most intuitive and concise representation [33]. We used it in some CML models for the generation process.
3.3 GT4SD Toolkit
The Generative Toolkit for Scientific Discovery (GT4SD) [28] handles molecules, calculates their properties, provides a variety of pretrained models and a framework to train CML models. We used six pretrained models, each of them generating a set of 2000 molecules. They were trained in the ChEMBL [13] dataset and made available by the GT4SD team. The first half of them uses SMILES: Variational Auto Encoder (VAE) [14], Objective Reinforced Generative Adversarial Networks (ORGAN) [26] and Adversarial Auto Encoders (AAE) [17]. The second half process the molecules as graphs: Molecular Scaffolds with Structural Motifs (MoLeR) [21], Graph Convolutional Policy Networks (GCPN) [34] and flow-based autoregressive model for molecular graph generation (GraphAF) [27].
3.4 Clus System
ClusFootnote 1 is a decision tree and rule induction system based on Predictive Clustering Trees. It unifies clustering and predictive modeling, and deals with complex prediction tasks such as multi-label classification. Within Clus, we used the hierarchical multi-label classifier Clus-HMC [30] to learn the intrinsic rules of the ChEBI taxonomy. We then used Clus-HMC to predict the taxonomy of our generated molecules based on their descriptors.
3.5 ChEBI Dataset
The Chemical Entities of Biological Interest (ChEBI) [8] is a dataset that contains information about molecules, such as SMILES representation and descriptors. It uses a Web Ontology Language (OWL) [1] to describe a taxonomy of molecules and their classification. Its information can be preprocessed and represented into a Directed Acyclic Graph (DAG) where each node represents a taxonomy label. It is also possible to prune the DAG, leaving the most important labels and molecules. One can also store the taxonomy information, obtain statistics, and compare groups of molecules.
The ChEBI DAG has three ontologies with nodes belonging to one or more of them, inheriting semantic meaning with their connections. The first ontology classifies molecular entities and chemical substances. The second expresses a molecular role, which is a particular behavior that a material may exhibit. The third expresses atomic particles smaller than an atom, e.g., neutrons and protons.
In order to improve the performance of the hierarchical classifier, we created a library to process and prune the original ChEBI DAG. After pruning, we kept only molecules with role information, since they are more relevant for drug discovery, and only nodes with at least 100 annotated molecules. With this, we ended up excluding the particle graph and many nodes from the structural and role graphs, obtaining a final DAG with 35032 molecules. Table 1 shows the statistics of the pruned DAG. The pruned DAG contains 72 nodes shared between the two ontologies, where 37 are leaves. The table also presents the leaves with most molecules classified for each graph. As the analysis focus on the role information, the most populated leaf by molecules in the role graph has significantly more molecules than the most populated leaf in the structural graph. The hierarchical classifier is trained in the pruned ChEBI DAG and used to predict the taxonomy of the molecules generated by the GT4SD toolkit. Thus, the classifier may infer role or structural information of the generated molecules.
3.6 Internal and External Diversity Measures
To compare molecules based on their structural information, the fingerprints representation can be used with the Tanimoto distance (Eq. 1) [2], where a and b are molecules. The internal diversity measures the variety of the molecules in a given set. It is defined in Eq. 2 as the mean of the sum of the Tanimoto distance applied to every pair of molecules in a set, where A is a set of molecules, and a and b are molecules from A. The external diversity is analogous, with the main difference of using two different sets to define the pairs of molecules. It is presented in Eq. 3, with a and b molecules from sets A and B respectively.
The external and internal diversities have a complexity problem. As the sets of molecules grow in size, the time required to calculate the measures grows proportionally to the square of the sizes of the sets [3]. As external diversity is used to compare groups of molecules, we used it as a guideline to propose a faster measure to compare groups of molecules: the hierarchical diversity.
3.7 Proposed Hierarchical Diversity Measure
Our proposed diversity uses taxonomies to compare different groups of molecules. A DAG is created for each set of molecules, and one is chosen as reference. Each node from the DAG stores how many molecules are classified in it, and the nodes are ordered constructing the first DAG distribution. The second DAG uses the same labels in the same order as the first DAG to construct its distribution in an analogous way. Finally, the distance of the distributions of molecules is measured using the Wasserstein Distance [29]. It is helpful to compare large groups of molecules since its complexity is way smaller than the external diversity. Equation 4 presents the Wassistein distance between distributions u and v. These are the distributions of the DAGs molecules over their nodes.
The hierarchical distance is asymmetric. Since we need to choose one of the DAGs as reference, the results may be different depending on the chosen DAG. However, our results in Sect. 4 showed these differences were not significant.
3.8 Methodology Overview
Figure 1 presents an overview of our methodology. The ChEBI dataset, in a OWL format (ChEBI.OWL), is preprocessed into a DAG, which is pruned and saved in two files: ChEBI.arff and ChEBI.obj. Each method from the GT4SD library then generates a Method.arff with molecules. Clus-HMC classifier is then trained with ChEBI.arff molecules, making predictions for the molecules in the Method.arff files. The predictions are stored in a Method.pred.arff file, and converted into the Method.obj format, which is used along with the ChEBI.obj to generate the results discussed in Sect. 4. All our implementations and datasets are freely availableFootnote 2.
4 Results and Discussions
This section presents an analysis of our generated molecules. For each of the six generators from the GT4SD toolkit, we generated a group of 2 thousand molecules. First, we analyze some of the molecules properties, and visualize molecules using their descriptors space. Next, we present the classification of the molecules by the Clus-HMC classifier. Internal and external diversities are then discussed, together with our proposed hierarchical distance.
For each molecule, 21 descriptors were calculated, including molecular weight, related to the number and type of the molecule atoms, PlogP, related to the solubility of the molecule in water, and all properties presented in Sect. 3.1. Table 2 presents information about some of the most important descriptors for each generated set of molecules. The sets generated by the AAE model differs from the other sets, having in general higher mean and variance for the molecular weight and PlogP descriptors. We can imply that many molecules in this set violate at least one rule of the RO5 rule. On the other hand, the set generated by GCPN showed the lowest values and variations for these two properties.
To visualize the multiple dimensions of the molecules, we applied two dimensionality reduction methods: Principal Component Analysis (PCA) [16] and Uniform Manifold Approximation and Projection (UMAP) [22]. PCA was applied in the a dataset with 12 thousand feature vectors (6 methods \(\times \) 2 thousand molecules), each one with 21 dimensions. PCA was also applied followed by UMAP, which rearranges data based on its proximity. This rearrangement may help to visualize overlapped data and clusters of data.
Figure 2 illustrates how the molecules are distributed in the PCA and UMAP spaces. The most outstanding group of molecules are the ones generated by AAE and GraphAF, while the other four groups appear to be very similar to each other. Although many of the points in the PCA space overlap, we can see where each group is located in relation to each other. In the UMAP space, the points hardly overlap, but it is hard to define a region for each group of molecules.
Clus-HMC was trained with a 10-fold cross-validation strategy using the 35032 molecules extracted from the ChEBI dataset. We used the Area Under Receiver Operating Characteristic (AUROC) and the Area Under Precision Recall Curve (AUPRC) to evaluate the classifier. AUROC measures the ability of a classifier to distinguish between positive and negative classes, while AUPRC summarizes the relationship between precision and recall. Table 3 shows the average AUPRC and AUROC values obtained. The values are considerably high, except for the simple average AUPRC which does not consider class imbalance.
We classified the molecules into classes of the ChEBI taxonomy. Tables 4 and 5 present information about the role and structural DAGs for the molecules generated by each method. Although we can provide the information individually for each molecule, we show the results for groups of molecules in order to have a general view of the properties generated by each method. We can observe that the ORGAN, VAE and MoLeR methods have close statistics, suggesting that their generated molecules are similar to each other. This same analysis can be done with GCPN and GraphAF. The molecules generated by AAE are the ones that differ the most from the other groups of molecules.
Table 6 shows some classes predicted by the hierarchical classifier for the generated molecules, and the corresponding classification percentages. The classes include biochemical functions, metabolites and subclasses of metabolites, alkaloids and an alkaloid subclass. With the hierarchical classification, we can reduce the number of molecules to search, allowing a more localized and focused search on certain classes or chemical characteristics. Some of the predicted classes have medicinal and pharmacological potential, such as Harmala alkaloids [20], and are present in pharmaceutical drug compounds, such as metabolites [11].
We used internal and external diversity to verify the similarities of the groups of molecules. The main diagonal of Table 7 shows the internal diversity, while the external diversity is shown in the other cells. Three main observations can be made: i) the GraphAF and GCPN molecules have the highest internal diversities; ii) the molecules generated by GraphAF and GCPN have the highest external diversities; and iii) the molecules generated by AAE and VAE have the smallest external diversities. These observations are useful to interpret the relations between the sets of molecules, and as a baseline to experiment if our proposed hierarchical distance is also capable to capture these relations.
Given the difficulty to visualize groups of molecules in Fig. 2, Fig. 3 shows only molecules generated by ORGAN and VAE. They were chosen to validate how our hierarchical distance evaluated two similar sets of molecules. Since the groups are close, a small value is expected. Figure 4 shows the distributions of the same molecules over the DAG nodes. Again, the distributions are very similar, and the hierarchical distance should be small.
Figure 5 shows the visualization of the descriptors of the molecules generated by AAE and GraphAF. They were chosen to validate how the hierarchical distance differs two dissimilar sets of molecules. They are not so close to each other, and a high hierarchical distance is expected. Figure 6 shows the distributions of the same molecules over the DAG nodes. We see a sensitive dissonance, which must result in a high hierarchical distance.
Table 8 shows our proposed hierarchical distance values for every pair of sets of molecules. From the results, we can separate the molecules in two clusters. The first one contains molecules generated by GraphAF and GCPN, showing relative small distance between them and high distances to the other groups. The second cluster is formed by the other molecules, with high similarity among them and high distances to the first cluster. The results of Table 8 are in consonance with the ones from Table 7, except that diversity values are constrained between 0 and 1. The hierarchical distance varies according to the DAG sizes and the dissonance between the sets of molecules. Also, in accordance to what we can see in Figs. 3, 4, 5 and 6, Table 8 shows small hierarchical distance differences between the groups of molecules generated by ORGAN and VAE, and high differences between the groups of molecules generated by AAE and GraphAF.
5 Conclusions and Future Works
In this paper we presented a study on Constructive Machine Learning applied to de novo drug-like molecules design. We generated molecules using six different methods from the GT4SD toolkit, and compared them with internal and external diversity measures. We proposed to use a hierarchical multi-label classifier to classify the molecules into classes of a known taxonomy, and compared groups of molecules based on their classification. For this comparison, we proposed a new hierarchical diversity measure that showed to be consistent in comparing groups of molecules based on their taxonomy information.
The ChEBI dataset was used to train a hierarchical multi-label classifier, since it contains taxonomy information from a variety of molecules, and is much smaller than other databases. Our proposal can be used to expand the ChEBI dataset by classifying new molecules in its taxonomy. It also can be used in drug discovery, classifying the generated molecules to narrow down the number of candidates, and selecting the molecules with specific chemical characteristics.
Once we classified groups of molecules into taxonomies, our proposed hierarchical distance obtained results similar to those from external diversity, but in a fraction of the time. External diversity has a limited range between 0 and 1, whereas our hierarchical distance can vary based on the size and diversity of the compared groups. The experiments showed that the hierarchical distance is more meaningful when calculated for similar sized groups of molecules.
As future works, the hierarchical distance could be extended to other contexts, to compare groups of different molecules that can be classified into a taxonomy. Also, we intend to investigate how the selected molecules can be used to fine tune their generators, in order to create a specific domain generation model.
References
Antoniou, G., Harmelen, F.v.: Web ontology language: owl. In: Handbook on ontologies, pp. 67–92. Springer (2004)
Bajusz, D., Rácz, A., Héberger, K.: Why is Tanimoto index an appropriate choice for fingerprint-based similarity calculations? J. Cheminform. 7 (2015)
Benhenda, M.: ChemGAN challenge for drug discovery: can AI reproduce natural chemical diversity? arXiv preprint arXiv:1708.08227 (2017)
Bickerton, G.R., Paolini, G.V., Besnard, J., Muresan, S., Hopkins, A.L.: Quantifying the chemical beauty of drugs. Nat. Chem. 4(2), 90–98 (2012)
Bjerrum, E.J., Threlfall, R.: Molecular generation with recurrent neural networks (RNNs). arXiv preprint arXiv:1705.04612 (2017)
Brown, N., Ertl, P., Lewis, R., Luksch, T., Reker, D., Schneider, N.: Artificial intelligence in chemistry and drug design. J. Comput. Aided Mol. Des. 34(7), 709–715 (2020). https://doi.org/10.1007/s10822-020-00317-x
Cao, D.S., Xu, Q., Hu, Q., Liang, Y.Z.: Manual for ChemoPy (2013)
Degtyarenko, K., et al.: ChEBI: a database and ontology for chemical entities of biological interest. Nucleic Acids Res. 36(suppl 1), D344–D350 (2007)
DiMasi, J.A., Grabowski, H.G., Hansen, R.W.: Innovation in the pharmaceutical industry: new estimates of r &d costs. J. Health Econ. 47, 20–33 (2016)
Elton, D.C., Boukouvalas, Z., Fuge, M.D., Chung, P.W.: Deep learning for molecular design-a review of the state of the art. Mol. Syst. Des. Eng. 4(4), 828–849 (2019)
Evans, L., Phipps, R., Shanu-Wilson, J., Steele, J., Wrigley, S.: Methods for metabolite generation and characterization by NMR. In: Ma, S., Chowdhury, S.K. (eds.) Identification and Quantification of Drugs, Metabolites, Drug Metabolizing Enzymes, and Transporters (Second Edition), pp. 119–150. Elsevier, Amsterdam, second edition. (2020)
Foster, D.: Generative Deep Learning: Teaching Machines to Paint, Write, Compose, and Play. O’Reilly Media (2019)
Gaulton, A., et al.: The ChEMBL database in 2017. Nucleic Acids Res. 45(D1), D945–D954 (2017)
Gómez-Bombarelli, R., et al.: Automatic chemical design using a data-driven continuous representation of molecules. ACS Cent. Sci. 4(2), 268–276 (2018)
Gupta, A., Müller, A.T., Huisman, B.J., Fuchs, J.A., Schneider, P., Schneider, G.: Generative recurrent networks for de novo drug design. Mol. Inf. 37(1–2), 1700111 (2018)
Jolliffe, I.: Principal component analysis. Encyclopedia of Statistics in Behavioral Science (2005)
Kadurin, A., Nikolenko, S., Khrabrov, K., Aliper, A., Zhavoronkov, A.: druGAN: an advanced generative adversarial autoencoder model for de novo generation of new molecules with desired molecular properties in silico. Mol. Pharm. 14(9), 3098–3104 (2017)
Kim, S., et al.: PubChem 2019 update: improved access to chemical data. Nucleic Acids Res. 47(D1), D1102–D1109 (2019)
Lipinski, C.A.: Lead-and drug-like compounds: the rule-of-five revolution. Drug Discov. Today Technol. 1(4), 337–341 (2004)
Marwat, S.K., ur Rehman, F.: Medicinal and pharmacological potential of harmala (peganum harmala l.) seeds. In: Preedy, V.R., Watson, R.R., Patel, V.B. (eds.) Nuts and Seeds in Health and Disease Prevention, pp. 585–599. Academic Press, San Diego (2011)
Maziarz, K., et al.: Learning to extend molecular scaffolds with structural motifs. arXiv preprint arXiv:2103.03864 (2021)
McInnes, L., Healy, J., Melville, J.: UMAP: uniform manifold approximation and projection for dimension reduction. arXiv preprint arXiv:1802.03426 (2018)
Mitchell, J.B.: Machine learning methods in chemoinformatics. Wiley Interdisc. Rev. Comput. Mol. Sci. 4(5), 468–481 (2014)
Olivecrona, M., Blaschke, T., Engkvist, O., Chen, H.: Molecular de-novo design through deep reinforcement learning. J. cheminform. 9(1), 48 (2017)
Papamakarios, G., Pavlakou, T., Murray, I.: Masked autoregressive flow for density estimation. In: Advances in Neural Information Processing Systems, vol. 30 (2017)
Sanchez-Lengeling, B., Outeiral, C., Guimaraes, G.L., Aspuru-Guzik, A.: Optimizing distributions over molecular space. an objective-reinforced generative adversarial network for inverse-design chemistry (organic). ChemRxiv (2017)
Shi, C., Xu, M., Zhu, Z., Zhang, W., Zhang, M., Tang, J.: GraphAF: a flow-based autoregressive model for molecular graph generation. arXiv preprint arXiv:2001.09382 (2020)
Team, G.: GT4SD (Generative Toolkit for Scientific Discovery) (2022)
Vallender, S.: Calculation of the Wasserstein distance between probability distributions on the line. Theor. Probab. Appl. 18(4), 784–786 (1974)
Vens, C., Struyf, J., Schietgat, L., Džeroski, S., Blockeel, H.: Decision trees for hierarchical multi-label classification. Mach. Learn. 73(2), 185–214 (2008)
Wegner, J.K., et al.: Cheminformatics. Commun. ACM 55(11), 65–75 (2012)
Weininger, D.: Smiles, a chemical language and information system. 1. introduction to methodology and encoding rules. J. Chem. Inf. Comput. Sci. 28(1), 31–36 (1988)
Xiong, J., Xiong, Z., Chen, K., Jiang, H., Zheng, M.: Graph neural networks for automated de novo drug design. Drug Discovery Today 26(6), 1382–1393 (2021)
You, J., Liu, B., Ying, Z., Pande, V., Leskovec, J.: Graph convolutional policy network for goal-directed molecular graph generation. In: Advances in neural information processing systems, vol. 31 (2018)
Acknowledgments
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001. The authors also thank the Brazilian research agencies FAPESP and CNPq for financial support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2023 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
de Souza Silva, R.R., Cerri, R. (2023). Constructive Machine Learning and Hierarchical Multi-label Classification for Molecules Design. In: Naldi, M.C., Bianchi, R.A.C. (eds) Intelligent Systems. BRACIS 2023. Lecture Notes in Computer Science(), vol 14196. Springer, Cham. https://doi.org/10.1007/978-3-031-45389-2_19
Download citation
DOI: https://doi.org/10.1007/978-3-031-45389-2_19
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-45388-5
Online ISBN: 978-3-031-45389-2
eBook Packages: Computer ScienceComputer Science (R0)