Abstract
A common approach for analyzing medical images on volumetric data is to employ deep 2D convolutional neural networks (2D CNN) on each individual slice of the volume. This is largely attributed to the challenges posed by the nature of three-dimensional data: variable volume size, high GPU and RAM requirements, costly parameter optimization, etc. However, handling the individual slices independently in 2D CNNs deliberately discards the temporal information that is contained within the depth of the volumes, which tends to result in poor performance. In order to maintain temporal information, current solutions reduce the temporal dimensionality of the data using non-adaptive sampling processes, such as taking slices at given intervals or by means of interpolation. However, although this allows to keep some temporal information, it may discard important slices. In this paper, we propose a method based on GradCam to select meaningful slices in computed tomography volumes by evaluating the activation map to reduce temporal data dimensionality. Extensive experiments demonstrate the effectiveness of our method when compared to the current state of the art.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
X-ray is a form of electromagnetic radiation, like visible light. It is less energetic than gamma rays and more energetic than ultraviolet light. While the human body is mostly opaque to visible light, X-rays easily pass through soft tissue, such as organs and muscles, but not as easily through hard tissue, such as bones and teeth. Consequently, X-ray imaging is well-suited for examining skeletal structures, but not so much for soft tissue, as it is the case of the brain. Computed tomography (CT) addresses this limitation, allowing thorough observation of internal structures, including the brain, the lungs, and other organs. The technique is non-invasive and provides good cross-sectional visualization. Additionally, CT exams are usually quick, with a single imaging session often completed in less than a few minutes.
CT scans are obtained from multiple shots at different angles, and with complex geometric transformations they produce volumes of data called slices. A set of slices is comparable to a collection of images that provide information on body sections. In a sense, CT scans are considered a type of three-dimensional data, as each slice can be transformed into a gray-scale image in a two-dimensional space and a full CT exam can be transformed into a sequence of images.
They are used by medical professionals to diagnose a multitude of diseases and conditions, and may provide a rich source of information to Artificial Intelligence (AI) methods. However, many AI engines are unable to make the best use of CT data. The reason is the often large volume of data, which renders the computational cost involved in training and inference infeasible for many information systems [1]. Instead, researchers often resort to two-dimensional approaches, which naturally disregard the sequential aspect of the volumetric data. While this reduces the computational cost and allows models to be trained and employed without the requirement of supercomputers, performance is usually also reduced. Furthermore, even when approaches designed to cope with three-dimensional (3D) data are used, such as 3D convolutional neural networks (CNN3D), performance tends to be lower than expected, especially when compared to two-dimensional approaches. This is primarily due to the difficulty of adjusting hyper-parameters, which require very time-consuming validation steps, and the necessity of reducing the resolution of the scans in favor of temporal data.
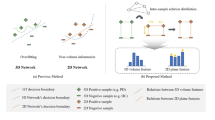
State-of-the-art approaches attempt to reduce the data’s temporal dimensionality while preserving image resolution as much as possible. However, most techniques rely on non-adaptive sampling processes, such as taking slices at given intervals or interpolating through slices. As a result, slices that are irrelevant to the application may be selected while important slices are discarded.
In this work, we address the problem of selecting slices that are more relevant for machine learning models, while simultaneously discarding slices containing less information about a given decision problem (e.g., detecting signs of a disease in volumetric CT). Our approach is based in the Grad-CAM technique [25]. Ideally, this should result in accurate models with shorter training times and lower hardware demands. As a case study, two datasets addressing two different decision problems are investigated in this paper: a set of lung CTs to determine if the patient is infected with the COVID-19 virus; and a dataset for intracranial hemorrhage detection. The highlights of our paper are:
-
We employ Gradient-weighted Class Activation Mapping (Grad-CAM) to eliminate the manual task of selecting meaningful slices.
-
We use depthwise convolutions [7] and Grad-CAM on volumetric CT data to preserve temporal order and facilitate relevant slice selection through adaptation of CNN [27].
-
Our proposed method reduces the temporal dimensionality of volumetric CT data, providing an advantage when a low amount of processing power is available.
-
The GSS method consistently shows the best results compared to other slice selection methods, both in terms of AUC and F1 Score. This is true for all configurations and for both deep learning models (C3D and 3DCNN-C) investigated in this paper. Therefore, the GSS method is the most promising for the task of selecting meaningful slices on CT volumes.
2 Related Work
Deep learning has been utilized in various medical domains [3, 5, 19, 24, 31]. Many of these studies concentrate on techniques developed for two-dimensional (2D) data, such as images. Since volumetric CTs are inherently 3D, one common approach is to analyze each CT slice individually using algorithms designed for 2D data, primarily state-of-the-art 2D CNNs [10, 15, 18, 23, 26, 28]. However, there is evidence suggesting that utilizing 3D data from CT scans leads to improved results [2, 14, 16, 17, 21], as it maintains the depth properties of the CT scans.
Several challenges arise when processing CT scans with volumetric data. Generally, data complexity increases exponentially with each added dimension [20]. In machine learning, this often results in substantially larger demands for memory, computation, and training data [12] because the complexity of the model must grow to more properly represent the complexity of the date. In the case of deep neural networks (DNN), learning from 3D data typically demands more layers and neurons than learning from 2D data. In other words, working with 3D data is significantly more computationally expensive, and one might not have enough resources to train with 3D data.
To provide some insight into this computational cost, we summarize the requirements for three machine learning methods employed in this study when classifying examples of the Mosmed data set. The Mosmed data set contains volumetric CT scan data that we resize in both width (image resolution) and depth (number of slices per instance). Our analysis is presented in Table 1. In all instances, the CT scan slice numbers were standardized to either 12 or 30, and the resolution of each scan remained at 512\(\,\times \,\)512 pixels or was scaled down to 224\(\,\times \,\)244 pixels. We measured the model size in memory, the number of trainable parameters, the number of floating-point operations (FLOPs), and the GPU memory needed to perform tasks with each model. The number of FLOPs directly influences the model’s execution time, and the required GPU memory determines whether the model can be used with the available computational resources. For example, SqueezeNet is a compact neural network designed for low-end devices, requiring minimal GPU memory. Conversely, the traditional 3D CNN architecture demands nearly 100 gigabytes of GPU memory to train and test 30 slices at full resolution. At the time of writing this article, this memory requirement exceeds the capacity of all consumer-grade devices on the market.
Typical CT exam equipments can produce volumes ranging from 2 to 640 slices. Selecting the most meaningful slices is the optimal approach for analyzing this data without discarding temporal information. Even if one possesses sufficient computational power to process all 640 slices, reducing data complexity is usually preferable, as less complex models tend to generalize better and decrease the risk of overfitting [4]. Slice selection can be achieved using adaptive or non-adaptive approaches. Adaptive approaches examine the slice content to determine which ones contain the most relevant information, while non-adaptive methods select subsets of slices regardless of their content. There are five primary techniques found in the literature, described in this section. The first three are examples of non-adaptive methods, the fourth is an adaptive approach, while the remaining one is also non-adaptive.
Discard by Slice Similarity (DSS) is a category of methods where a similarity-based algorithm (e.g., Structural Similarity Index Measure (SSIM), Mean Squared Error (MSE), Euclidean Distance) compares each slice to its successor. Based on a defined threshold, the most similar slices are eliminated [29]. This approach aims to reduce redundancy in the data while preserving unique and informative slices.
Subset Slice Selection (SSS) is a method proposed in [34]. In this approach, slices are sampled from three specific positions of a volumetric CT: start, middle, and end. To achieve this, the volume is first divided into three equal parts. Then, a desired number of slices are extracted from each part. This method aims to capture the representative information from different regions of the CT volume.
Even Slice Selection (ESS), also known as Uniform Slice Selection, is a technique commonly used in video processing. In ESS, a “spacing factor” is calculated to enable the selection of equidistant slices [33]. Given a volume with D slices and a desired number of \(K << D\) slices, ESS works by dividing the volume into disjoint subsets of approximately \(\frac{D}{K}\) slices. Then, only the first slice of each subset is retained. In contrast to SSS, this technique mitigates semantic losses of temporal data. Algorithm 1 summarizes the ESS method.

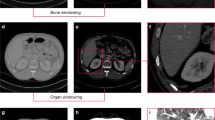
Slice Selection by Object Detection (SSOD) consists of methods in which an algorithm scans each slice and determines whether an object of interest is present. For instance, an AI model may be employed on CT scans to detect which slices display segments of organs such as lungs, brain, or other targeted organs [8]. The slices displaying the desired organ are then selected, while the others are discarded. While this method adapts to the data, it overlooks the inherent temporal relationships between slices. Often, it discards the initial and final subsets of slices, as the largest portion of the target organ is typically located in the middle of the volume.
As previously mentioned, SSOD is an adaptive method, whereas DSS, SSS, and ESS are non-adaptive. In some applications, employing non-adaptive strategies to discard slices may lead to loss of essential information (e.g., a critical segment of an organ may not be present in any of the selected slices). Spline Interpolation Zoom (SIZ) is a non-adaptive strategy aimed at reducing these losses. In SIZ, the temporal dimension is reduced by interpolating all slices. The volume is enlarged or compressed by replicating the closest pixel of each slice along the depth/z-axis. Although each resulting slice may be less precisely “located” in time, this technique may be advantageous for various applications since it does not produce “gaps” in the temporal axis [13, 32]. As summarized in Algorithm 2, given K as the desired number of slices and D as the total number of slices of a volume, zoom is performed along the z-axis by a factor of \(\frac{1}{D/K}\) using interpolation by splines [9].

Our work primarily focuses on comparing non-adaptive techniques against our proposed method because, although our method is adaptive in nature, many of the available adaptive methods, including SSOD, do not leverage the temporal relationship present in the slices when selecting. This omission may result in a loss of critical information. Furthermore, most adaptive slice selection methods in the medical domain currently focus on object detection rather than preserving temporal information. We believe our adaptive approach that emphasizes temporal relationships offers a unique perspective in the slice selection domain, warranting its comparison with more traditional non-adaptive methods.
3 Materials and Methods
In this section we consider the definitions provided by Tran et al. [30], a primary reference for 3D learning, to help us to explain our proposed method. In that work, Tran et al. provide empirical evidence that 3D convolutions are effective feature extractors when modelling appearance and motion or depth simultaneously. This makes them well-suited for learning sequence and spatiotemporal data. For instance, in 3D CNNs, even the output of each layer is a volume.
In 2D convolution, the kernel is a 3-dimensional matrix and both the input layer and the filters have the same depth (channel number equals their kernel number). However, the 3D filter only moves in two directions, as in over the height and the width of the image. In other words, it deals with only the spatial dimensions of the input. Therefore, even when using 3D filters, the output is a 2D array.
Conversely, 3D convolution not only involves a 3D filter, but the filter can move in all three directions—height, width and time/sequence. At each position, multiplication and elementary addition provide a number. As the filter slides through the 3D space, the output numbers are also arranged in a 3D space, i.e. the output is 3D data. CNNs exploit the spatially-local correlation by enforcing a local connectivity pattern between neurons of adjacent layers. Such 3D convolutions also take temporal features into account, typically through a sliding window, which is a filter with trainable weights over the input, and producing as output a weighted sum of weights and input. The weighted sum is the feature space used as the input for the next layers.
In this work we employ depthwise convolution, which is a type of convolution where a single convolutional filter is applied for each input channel. In regular 2D convolution performed over multiple input channels, the filter is as deep as the input and mixes channels to generate each element of the output feature map. In contrast, depthwise convolution keeps each channel separated. In our case, it maintains each slice separated, preserving the temporal relationship at different timestamps. We summarize the steps involved in this process bellow:
-
1.
The input tensor of 3 dimensions (volumetric CT, in our case) is divided into separate channels/slices.
-
2.
Each slice is convolved with its respective filter.
-
3.
The obtained convolved output are stacked together to provide the output as an entire 3D tensor.
Figure 1 exemplifies this process. At the top part of the image an instance containing three slices (a volumetric CT) is shown. Each slice is trated separated and convolution is only performed within each slice. It is necessary to maintain the depth size until the final convolution so that the feature maps of each slice is preserved to allow the extraction of Grad-Cam heatmap, which is performed in our proposed method, as detailed in next sections.
3.1 Inference Method
The proposed method consists of a 3DCNN designed to capture the activation map of a volumetric CT. The objective is to obtain as output a 3D matrix representing the regions of interest in each slice of the volume. This is achieved by adapting the Grad-Cam technique, which uses the gradients obtained when evaluating the class of interest by flowing to the last convolutional layer in order to map the activation regions of an image. Grad-CAM requires no re-training and is broadly applicable to any CNN-based architectures. Additionally, the generated heatmaps may provide explanations, making the predictions understandable to users, consequently helping the users to trust the predictions made by the system, especially in medical applications.
As previously mentioned, we use a model that preserves the depth dimension up to the last convolutional layer, performing depthwise convolutions on the volume so as to maintain the order of each slice. In this way, a feature map is created for each slice. However, since CT are temporal high-dimensional data, a critical point here is the viability of training a 3DCNN model using all the slices of the volume, depending on the chosen hardware. Therefore, in order to allow this training, we focus on providing a condition of lower computational cost to train the model by applying image compression techniques to identify macro-regions of interest. Thus, the resizing of the width and height dimensions is a fundamental aspect.
The following steps were performed to obtain our proposed Grad-CAM Slice Selection method (GSS), in which Grad-Cam is applied to obtain the meaningful slices:
-
1.
Normalize the volumetric data with one of the techniques described in Sect. 2. SIZ (Sect. 2) was used in this work.
-
2.
Train a CNN3D in the classification problem represented by the volumetric data preserving the depth dimension, e.g. the binary classification of data as COVID-19 or not.
-
3.
Remove the layers stacked after the last convolutional layer after finishing the network training, since they are no longer needed.
-
4.
Extract the representative matrix of heatmaps.
These steps are also summarized in Algorithm 3 and are further visualized in Fig. 2 that shows that the last Conv Layer will be analyzed for the heatmap acquisition.

4 Experimental Results
In this section, we present the experimental results obtained using two 3DCNN architectures: C3D and 3DCNN-C. The experiments were conducted using the canonic houldout partitions established as training and test splits of the two datasets investigated in order to provide state-of-the-art comparison. The datasets employed are MosmedData [22] and CQ500 [6], detailed below. One notable aspect of our experimental design is the conscious decision to not employ any data augmentation techniques. Data augmentation is often employed to artificially expand the dataset size by introducing variations such as rotation, flipping, and scaling. While this can improve the model’s generalization ability, it also introduces an additional layer of complexity that could potentially skew results or make direct comparisons with other methodologies more challenging. In our study, we aim to evaluate the efficacy of the deep learning architectures in recognizing patterns in unaltered, real-world medical imaging data. By excluding data augmentation, we seek to establish a baseline performance metric that reflects the model capabilities to generalize from the raw, original data, thereby ensuring that any observed performance differences are attributable solely to the architectures themselves, rather than to external manipulations of the data.
MosMedData contains anonymized human lung CT scans with COVID-19 related findings, as well as those without such findings. The CT scans were obtained between 1st March 2020 and 25th April 2020, provided by medical hospitals in Moscow, Russia.
CQ500 is a dataset that contains 491 scans with 193,317 slices, provided as anonymized DICOMs by the Centre for Advanced Research in Imaging, Neurosciences, and Genomics (CARING), New Delhi, India. The reads were performed by three radiologists with 8, 12, and 20 years of experience in cranial CT interpretation, respectively.
The 3D data from these datasets are in NIfTI format, which are volumetric (3D) images. After pre-processing, all images were set to a size of 512 with variable depth. Due to the nature of CT datasets, the images were imported as color Grayscale images, demanding modifications according to each network’s input for data compatibility. In addition, the datasets were partitioned following the holdout procedure into 70-15-15 for training, test and validations subsets, respectively.
The two 3DCNN models investigated were trained end-to-end on both datasets for 100 epochs with early stopping method so as to better tackle potential overfitting. Moreover, overfitting was also addressed by incorporating dropout and L1/L2 regularization techniques. The Adam optimizer was used in both architectures with a learning rate of \(10^{-4}\), determined experimentally. Weights were initialized using Glorot Normal Initialization [11], with convolution layers featuring the activation function Rectified Linear Activation Function (ReLU) and the final layer using the activation function sigmoid for the binary problem. Training was performed in an environment with a Tesla V100-SXM2 video card with 16 Gb memory.
As previously mentioned, all four non-adaptive slice selection methods described in Sect. 2 are used in this work as baselines. The experiments were divided into two comparison scenarios according to the number of selected slices, precisely 30 and 12 slices were selected with each method and compared against the proposed GSS. Table 2 presents the results of this comparison on the two distinct datasets, Mosmed and CQ500, representing binary classification problems. Besides the number of selected frames (30 and 12), the four investigated slice selection methods (SSS, ESS, SIZ, and GSS) are also compared considering two different image resolutions: 512\(\,\times \,\)512 and 224\(\,\times \,\)224. Finally, two 3DCNN models are used: C3D and 3DCNN-C adapted from [27]. The evaluation metrics used are AUC (area under the ROC curve) and F1 Score.
Analyzing the results, the following conclusions are observed:
-
GSS consistently presents the best results compared to other methods, both in terms of AUC and F1 Score. This is true across all configurations and for both deep learning models (C3D and 3DCNN-C). Therefore, GSS is the most promising method for selecting the most relevant slices from tomography volumes.
-
The baseline SIZ outperforms SSS and ESS in general, but still cannot surpass the results from GSS.
-
The results show that selecting 30 frames generally performs better than selecting 12 frames. This indicates that having more frames available for the model can lead to better model performance.
-
The 512\(\,\times \,\)512 resolution generally produces better results compared to the 224\(\,\times \,\)224 resolution, suggesting that a higher resolution provides more valuable information for the model.
-
The 3DCNN-C model adapted from [27] tends to achieve better results than the C3D model across all configurations and datasets.
Selecting slices without decreasing accuracy is a significant advancement towards using AI as an assistant in clinical management with lower computational power requirements. Therefore, the development of a clinical prognostic model based on our AI system, utilizing CT parameters and clinical data, marks an essential step forward in using AI to assist clinical management in this kind of scenario.
5 Conclusions
In this paper we propose GSS (Grad-CAM Slice Selection), a technique to deal with the problem of classifying complex 3D spatiotemporal data in the form of volumetric computer tomography (CT) imagery. By selecting slices from each CT in a way that takes into account the classification problem at hand, we are able to dynamically select the slices that provide more information, and reduce the computational cost required to train and apply a model.
We compared different deep learning techniques under the same conditions when applied to classification problems with two well-known data sets. By applying the proposed GSS to each deep learning model before training and inference, we achieved accuracies on par with or surpassing the state-of-the-art and, in most cases, better than other agnostic sampling techniques. This was especially true for edge cases requiring a greater number of slices to be selected.
Our findings emphatically underline the superiority of the GSS approach combined with 30 frames at 512\(\,\times \,\)512 resolution, leveraging the CNN model cited in [27]. As anticipated, this high-resolution configuration with an optimal number of frames yielded the most promising results in the domain of tomography volume anomaly detection. However, the strategic selection between resolution and the number of frames remains pivotal. For detecting nuanced or minute pathologies, it is imperative to lean towards higher spatial resolutions. Conversely, for challenges deeply rooted in the three-dimensional intricacies, a richer frame selection is recommended.
These findings support the hypothesis that using Grad-CAM in deep learning models for learning simple and macro-relevant features in a CT volume dataset is effective and warrants further investigation. Subsequently, transferring this knowledge to another model to learn complex patterns proves fruitful.
References
Ahmed, E., et al.: A survey on deep learning advances on different 3D data representations. arXiv preprint arXiv:1808.01462 (2018)
Alebiosu, D.O., Dharmaratne, A., Lim, C.H.: Improving tuberculosis severity assessment in computed tomography images using novel DAvoU-net segmentation and deep learning framework. Expert Syst. Appl. 213, 119287 (2023)
Becker, A.S., et al.: Detection of tuberculosis patterns in digital photographs of chest X-Ray images using deep learning: feasibility study. Int. J. Tuberculosis Lung Disease 22(3), 328–335 (2018)
Bejani, M.M., Ghatee, M.: A systematic review on overfitting control in shallow and deep neural networks. Artif. Intell. Rev. 54(8), 6391–6438 (2021). https://doi.org/10.1007/s10462-021-09975-1
Chen, W.W., et al.: A deep learning approach to classify fabry cardiomyopathy from hypertrophic cardiomyopathy using cine imaging on cardiac magnetic resonance. Int. J. Biomed. Imaging 2024(1), 6114826 (2024)
Chilamkurthy, S., et al.: Development and validation of deep learning algorithms for detection of critical findings in head CT scans. arXiv preprint arXiv:1803.05854 (2018)
Chollet, F.: Xception: deep learning with depthwise separable convolutions. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 1251–1258 (2017)
da Cruz, L.B., et al.: Kidney segmentation from computed tomography images using deep neural network. Comput. Biol. Med. 123, 103906 (2020)
De Boor, C.: Bicubic spline interpolation. J. Math. Phys. 41(1–4), 212–218 (1962)
Gao, X.W., Hui, R., Tian, Z.: Classification of CT brain images based on deep learning networks. Comput. Methods Programs Biomed. 138, 49–56 (2017)
Glorot, X., Bengio, Y.: Understanding the difficulty of training deep feedforward neural networks. In: Proceedings of the Thirteenth International Conference on Artificial Intelligence and Statistics, pp. 249–256. JMLR Workshop and Conference Proceedings (2010)
Goodfellow, I., Bengio, Y., Courville, A.: Deep learning. MIT press (2016)
Gordaliza, P.M., Vaquero, J.J., Sharpe, S., Gleeson, F., Munoz-Barrutia, A.: A multi-task self-normalizing 3D-CNN to infer tuberculosis radiological manifestations. arXiv preprint arXiv:1907.12331 (2019)
Grewal, M., Srivastava, M.M., Kumar, P., Varadarajan, S.: RadNet: radiologist level accuracy using deep learning for hemorrhage detection in CT scans. In: 2018 IEEE 15th International Symposium on Biomedical Imaging (ISBI 2018), pp. 281–284. IEEE (2018)
Hamadi, A., Cheikh, N.B., Zouatine, Y., Menad, S.M.B., Djebbara, M.R.: ImageCLEF 2019: deep learning for tuberculosis CT image analysis. In: CLEF (Working Notes) (2019)
Huang, X., Shan, J., Vaidya, V.: Lung nodule detection in CT using 3D convolutional neural networks. In: 2017 IEEE 14th International Symposium on Biomedical Imaging (ISBI 2017), pp. 379–383. IEEE (2017)
Ji, S., Xu, W., Yang, M., Yu, K.: 3D convolutional neural networks for human action recognition. IEEE Trans. Pattern Anal. Mach. Intell. 35(1), 221–231 (2012)
Kavitha, S., Nandhinee, P., Harshana, S., S, J.S., Harrinei, K.: ImageCLEF 2019: a 2D convolutional neural network approach for severity scoring of lung tuberculosis using CT images. In: CLEF (Working Notes) (2019)
Lakhani, P., Sundaram, B.: Deep learning at chest radiography: automated classification of pulmonary tuberculosis by using convolutional neural networks. Radiology 284(2), 574–582 (2017)
LeCun, Y., Bengio, Y., Hinton, G.: Deep learning. Nature 521(7553), 436–444 (2015)
Li, B., Zhang, T., Xia, T.: Vehicle detection from 3D lidar using fully convolutional network. arXiv preprint arXiv:1608.07916 (2016)
Morozov, S.P., et al.: MosMedData: data set of 1110 chest CT scans performed during the Covid-19 epidemic. Digit. Diagn. 1(1), 49–59 (2020)
Ronneberger, O., Fischer, P., Brox, T.: U-Net: convolutional networks for biomedical image segmentation. In: International Conference on Medical Image Computing and Computer-Assisted Intervention, pp. 234–241. Springer (2015). https://doi.org/10.1007/978-3-319-24574-4_28
Santosh, K., Allu, S., Rajaraman, S., Antani, S.: Advances in deep learning for tuberculosis screening using chest X-rays: the last 5 years review. J. Med. Syst. 46(11), 82 (2022)
Selvaraju, R.R., Cogswell, M., Das, A., Vedantam, R., Parikh, D., Batra, D.: Grad-cam: visual explanations from deep networks via gradient-based localization. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 618–626 (2017)
Shah, A.A., Malik, H.A.M., Muhammad, A., Alourani, A., Butt, Z.A.: Deep learning ensemble 2D CNN approach towards the detection of lung cancer. Sci. Rep. 13(1), 2987 (2023)
da Silva, L.A., et al.: Spatio-temporal deep learning-based methods for defect detection: an industrial application study case. Appl. Sci. 11(22), 10861 (2021)
Sultana, A., et al.: A real time method for distinguishing COVID-19 utilizing 2D-CNN and transfer learning. Sensors 23(9), 4458 (2023)
Thasneem, A.H., Sathik, M.M., Mehaboobathunnisa, R.: A fast segmentation and efficient slice reconstruction technique for head CT images. J. Intell. Syst. 28(4), 533–547 (2019)
Tran, D., Bourdev, L., Fergus, R., Torresani, L., Paluri, M.: Learning spatiotemporal features with 3D convolutional networks. In: Proceedings of the IEEE International Conference on Computer Vision, pp. 4489–4497 (2015)
Wajgi, R., et al.: Optimized tuberculosis classification system for chest X-ray images: fusing hyperparameter tuning with transfer learning approaches. Engineering Reports, e12906 (2024)
Yang, J., et al.: Reinventing 2D convolutions for 3d images. IEEE J. Biomedical Health Inform. (2021)
Zhu, H., Liu, X., Mao, X., Wong, T.T.: Real-time deep video deinterlacing. arXiv preprint arXiv:1708.00187 (2017)
Zunair, H., Rahman, A., Mohammed, N., Cohen, J.P.: Uniformizing techniques to process CT scans with 3D CNNs for tuberculosis prediction. In: Rekik, I., Adeli, E., Park, S.H., Valdés Hernández, M.C. (eds.) Predictive Intelligence in Medicine: Third International Workshop, PRIME 2020, Held in Conjunction with MICCAI 2020, Lima, Peru, October 8, 2020, Proceedings, pp. 156–168. Springer International Publishing, Cham (2020). https://doi.org/10.1007/978-3-030-59354-4_15
Acknowledgments
This study was financed in part by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES-PROEX) - Finance Code 001. This work was partially supported by Amazonas State Research Support Foundation - FAPEAM - through the project PDPG/CAPES.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2025 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Silva, L.A.d., Santos, E.M.d., Giusti, R. (2025). Deep Learning Approach to Temporal Dimensionality Reduction of Volumetric Computed Tomography. In: Paes, A., Verri, F.A.N. (eds) Intelligent Systems. BRACIS 2024. Lecture Notes in Computer Science(), vol 15412. Springer, Cham. https://doi.org/10.1007/978-3-031-79029-4_21
Download citation
DOI: https://doi.org/10.1007/978-3-031-79029-4_21
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-79028-7
Online ISBN: 978-3-031-79029-4
eBook Packages: Computer ScienceComputer Science (R0)