Abstract
Diseases of the gastrointestinal (GI) tract are among the most common pathologies in the world population and are responsible for thousands of deaths every year. This work proposes an automatic method for detecting regions with GI Tract abnormalities, intending to reduce the number of lesions missed in Wireless Capsule Endoscopy (WCE) video exams by expert endoscopists. By taking advantage of convolutional neural networks (CNNs) and YOLO detection models, the proposed method not only increases the reliability of pathological detection in WCE images, but also sets a new benchmark in this field. Our results for binary classification between healthy and pathological images are promising, with an accuracy of 87.8%, precision of 91.6%, recall of 89.1% and F1-Score of 90.3%. In addition, the detection model showed an Intersection over Union (IoU) of 31.33% among all the images classified as pathological. The impact of this research is significant, as it provides a method capable of detecting GI Tract diseases in WCE images and contributing to better clinical decision-making and patient care.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
The gastrointestinal (GI) tract is a tubular system whose main task is digestion. Its structure is made up of the mouth, pharynx, esophagus, stomach, small intestine, large intestine, rectum and anal canal. Inflammatory diseases associated with the GI Tract have a high incidence rate in the world population, especially in developing countries, where colon, liver, and stomach cancer are among the most recurrent diseases [15]. According to the World Health Organization (WHO), in 2022, there were approximately 5.1 million cases of GI Tract cancers worldwide, and, in the Americas region alone, 431,116 deaths were recorded [27].
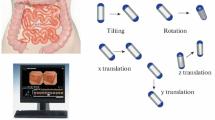
Based on the need to combat the harmful effects of GI Tract pathologies, endoscopy is one of the most widely used techniques for analyzing GI Tract pathologies, but it is characterized by being a painful and invasive process [12]. In contrast, videos can be taken using Wireless Capsule Endoscopy (WCE), a less aggressive and uncomfortable method. During this procedure, the patient swallows a capsule with a light source, a micro-camera, and a signal emitter that travels painlessly throughout the GI Tract until it is naturally expelled by the patient’s body. However, as they scan the entire GI Tract, the videos acquired by the capsule are around 8 h long. Therefore, analyzing this video is an arduous and tedious task, making it possible to miss abnormalities during the examination due to its dependency on the analyst’s constant concentration on the images [4]. For this reason, it is important to use automation to help the specialist analyze the video.
From this perspective, machine learning techniques, which consist of using a dataset to develop a computer-aided diagnosis (CAD) system, have been used in the analysis of medical images since the 1990s [13]. These techniques are widely used in medical tasks due to their ability to automatically learn discriminative features from raw data and to reduce the need for pre-processing and feature extraction [14]. An example of this type of method is Convolutional Neural Networks (CNNs), a computational resource inspired by the workings of the human brain commonly used in the processing and analysis of digital images [26]. Another well-known method is YOLO (You Only Look Once), which stands out as a real-time object detection algorithm and a single-stage detector that can predict all objects in a single pass through the image. In this way, the adoption of YOLO can help in the early detection and diagnosis of GI Tract diseases [19].
Furthermore, the importance of identifying the region of the pathology lies in enabling a more precise diagnosis, potentially avoiding additional invasive diagnostic procedures [12]. It allows for the comparison of images from subsequent exams to evaluate disease progression or treatment response and informing patients about the exact location of their condition can improve their understanding of the disease and increase treatment adherence. Thus, it becomes crucial to enhance the capability of tools to automatically detect and characterize pathologies, increasing the efficiency of CADs.
Therefore, this work aims to introduce detection methodologies for WCE images of the GI Tract, providing a solid foundation for subsequent research efforts to build upon the automated detection of diseases in WCE images. Thus, this study developed the application of a method to classify and detect the pathologies using the best CNN architecture experimented and YOLO. Next, Sect. 2 highlights the related work, Sect. 3 describes the proposed method, and Sect. 4 presents the results, experiments, comparisons, and discussions of the proposed method.
2 Related Works
In recent years, neural networks based on deep learning methods have proven to be a powerful tool for image analysis, including medical imaging tasks [3]. This has led to several advances in the field, including computer-aided classification and localization of pathologies from the GI Tract. This section presents some of the work that has been done on this subject.
Xu et al. (2019) introduced a multi-task anatomy detection convolutional neural network (MT-AD-CNN) designed to evaluate esophagogastroduodenoscopy inspection quality. The model classifies images and distinguishes between informative and non-informative frames, thus displaying detection boxes only on informative frames and reducing false positives. The MT-AD-CNN achieves a mean average precision (mAP) of 93.74% for the detection task and an accuracy of 98.77% for the classification task.
The work by Oukdach et al. (2024) presented a new framework that synergizes the strengths of CNNs and Vision Transformers (ViTs). This approach incorporates an attention mechanism within a CNN to extract local features. Experiments performed on the Kvasir Capsule, a large-scale dataset of WCE images [21], showed a promising result of 97% in accuracy, precision, recall, and F1-Score.
Muruganantham and Balakrishnan (2022) proposed a method for classifying images with ulcers, bleeding, polyps, or no abnormality using ResNet-50 associated with a self-attention mechanism aggregate spatial characteristics globally. The method was also tested on the Kvasir-Capsule dataset and achieved a classification accuracy of 94.7%.
In addition, Srivastava et al. (2022) proposed a focal modulation network called FocalConvNet. FocalConvNet makes use of focal modulation and an added convolutional block to extract features in order to improve detection. The study used 13 classes from anatomy and luminal findings from the Kvasir-Capsule dataset, obtaining F1-Score, recall, and MCC (Matthews Correlation Coefficient) of 67.34%, 63.73%, and 29.74, respectively.
As videos are sequences of captured images and the mentioned methods are applied per image (individual video frame), some work [5, 7, 17, 18, 22] do not clearly specify how sampling is done to split the training and test sets. Ideally, all video frames for a patient should be in the same set. Applying a random sampling of video frames can result in “data leakage” [9], as similar frames (neighbors) from the same patient can end up in both the training and test sets, compromising the results’ reliability.
In addition, although there are many published studies to assist in the diagnosis of GI pathologies in WCE images, there is a notable lack of studies focused on the detection of the region in which these anomalies are found. To this end, a pipeline was developed to classify abnormal images using the best-performing CNN architecture in the conducted experiments, followed by the detection of the region with the presence of the disease using YOLOv8 [8], a recent model for detecting objects from images in real-time. This work’s contribution lies in the proposal of an automated method that not only identifies the presence of different types of anomalies in WCE images but also indicates the region affected by the pathology, reducing the time and work involved in analyzing videos of WCE exams.
3 Materials and Method
This section describes the proposed method. Firstly, the image dataset used to train and evaluate the proposed method is determined, and then the best performing CNN architecture is selected to classify the images in the dataset. Then, the model that has shown the best performance during the experiments sends the images classified as abnormal for detection. Finally, the detection models trained for a specific class by YOLOv8, make predictions based on the images received. The predicted region is the one with the highest confidence between the model’s predictions. A representation of the process can be seen in Fig. 1.
3.1 Dataset
In this work, the experiments were carried out using the Kvasir-Capsule dataset, which was built by collecting WCE scans from a Norwegian hospital [21]. Currently, Kvasir-Capsule is the largest and most diverse WCE dataset with expert-labeled data and publicly available. It comprises a total of 117 videos collected from WCE exams, totaling around 4.7 million frames extracted from these videos.
In addition, the dataset features 47.238 frames labeled by specialists, 336\(\times \)336 in size and divided into two categories: anatomical findings, consisting of Pylorus, Ileocecal Valve and Ampulla of Vater, and luminal findings, composed of Angiectasia, Blood - fresh, Blood - hematin, Erosion, Erythema, Foreign Body, Polyp, Lymphangiectasia, Reduced Mucosal View and Ulcer, in addition to the Normal Clear Mucosa class, which represents normal images. Figure 2 shows the sample image for each class. The division of the dataset, the number of patients, and the availability of the bounding boxes per class can be seen in Table 1.
3.2 Classification Stage
In this stage, we previously implemented and evaluated CNN architectures to choose the model to be used in the method, defined the metrics to be used to evaluate the binary classification models, and trained and tested the dataset with the architecture that obtained the best performance.
Selection of CNN Model. After defining the dataset, the best-performing CNN is selected to classify the images. For this task, CNNs widely used in the literature in the field of image classification were chosen. The architectures evaluated in this stage were VGG [20], Resnet [6], Inception [23], Xception [1], EfficientNet [24] and EfficientNetV2 [25]. The architectures mentioned were used according to their respective original authors and the implementation provided by the Keras library [2].
Binary Classification Training and Evaluation. After choosing the best architecture to perform the binary classification, the model is evaluated during and after the training process using the following metrics:
where TP and TN represent correctly classified images with pathology and healthy images. In addition, FP represents the healthy images wrongly classified as having abnormalities, and FN means the classifications of pathological images as healthy images.
In the context of the proposed method’s objective, accuracy is useful for giving an overview of the model’s performance, but it is misleading in unbalanced datasets, as in the case of Kvasir-Capsule, where one class is more frequent than the other. Thus, precision is also used to ensure that images classified as containing pathologies actually do have pathologies, thus reducing false positives that can overload the region detection stage.
In addition, recall is crucial to ensure that as many images with pathologies as possible are identified, minimizing false negatives that could result in undetected pathologies. The F1-Score will be the most important metric to evaluate due to its balance between precision and recall, providing a single performance measure that considers both false positives and false negatives. Furthermore, although specificity is a useful metric in many contexts, in this work, the main focus is ensuring that all images with pathologies are identified (maximizing recall) and that the identified images contain pathologies (maximizing precision).
Thus, using the metrics presented, the model selects the images with pathologies and sends them to the detection phase to determine the region where the classified pathology appears.
3.3 Detection Stage
During this stage, the detection models based on YOLOv8 were trained and the pathological area was predicted based on the model with the highest prediction confidence.
YOLOv8 Architecture. YOLOv8 was unveiled in January 2023 by Ultralytics, providing five versions: YOLOv8n (nano), YOLOv8s (small), YOLOv8m (medium), YOLOv8l (large), and YOLOv8x (extra-large) [8]. YOLOv8 provides support for classification, segmentation, and object detection tasks. In this work, YOLOv8s was used as a model to detect the regions where anomalies appear in the dataset images. The YOLOv8 architecture can be seen in Fig. 3.
YOLOv8 architecture [8]
Among the differences in YOLOv8 compared to previous versions is the C2f (partial inter-stage bottleneck with two convolutions) module, which combines high-level features to improve detection accuracy based on contextual information. In addition, YOLOv8 has an approach that allows each branch to be focused on its task in order to improve the overall accuracy of the model. In the output layer of YOLOv8, the sigmoid function is used as the activation function to represent the probability of a bounding box containing an object. In addition, the softmax function is used for class probabilities.
YOLOv8 uses the CIoU (Complete Intersection over Union) and DFL (Distribution Focal Loss) loss functions for bounding box loss and binary cross-entropy for classification loss. These losses improved object detection performance, especially when dealing with smaller objects [11]. YOLOv8’s architecture uses a modified backbone CSPDarknet53 combined with an SPPF layer (Spatial Pyramid Pooling Fast), which speeds up the calculation by grouping features into a fixed-size map. Furthermore, each convolution has batch normalization and SiLU activation (Sigmoid Linear Unit), and the head of the architecture is decoupled, so the objectivity, classification, and regression tasks are processed independently [16].
Detection Model Training. After classifying the images with pathologies, the next step is to train the detection models with YOLOv8. The aim was to train a dedicated model to detect each class so that each model could predict based on the abnormal images received from the previous stage. YOLOv8s was therefore used to train the detection models. Along with accuracy and sensitivity, the following metrics were also used to measure the assertiveness of the predicted regions in relation to the true region with the pathology:
-
Average Precision (AP): \(\int _{0}^{1} P(R) \, dR \)
-
mean Average Precision (mAP): \((\frac{1}{n} \sum _{i=1}^{n} \text {AP}_i )\)
-
Intersection over Union (IoU): \(\frac{\text {Intersection Area}}{\text {Union area}} \)
mAP is a widely used metric in object detection tasks, as it helps calculate average accuracy at various IoU thresholds. Thus, it provides a comprehensive assessment of the algorithm’s accuracy in object detection. The mAP calculation considers the AP, the average of the accuracies calculated at various recovery points. The recovery point is where the rate of true positives is calculated in relation to the total number of positive examples.
Additionally, the IoU metric was used to measure how close the predicted bounding boxes are to the real box described in the dataset. This metric is calculated as the area of intersection over the area of the union of the prediction with the true area. It is commonly used to assess the accuracy of object detection. Thus, after training the dedicated models for each class, all these metrics were calculated and applied to a test set to evaluate the performance of each model.
Pathology Region Detection. After training the dedicated model for each class, the images classified as abnormal are passed through all the detection models to assess the confidence in the region predicted in each image. From the results stored in text files in the YOLO format, we will have access to the following prediction information: detected class, x and y coordinates of the center of the bounding box, width, and height of the predicted bounding box, and the prediction confidence. So this step consists of identifying the prediction with the highest confidence and then collecting the coordinates of the predicted bounding box.
Then, the given coordinates are translated into Cartesian format on top of the original image size and the region of intersection between the prediction and the true location of the pathology is stipulated. Once all the images have been passed through and the region of interest has been identified, the IoU corresponding to all the intersection regions is calculated in order to obtain an overall evaluation of the proposed method.
4 Results and Discussion
This section presents the results achieved using the proposed method for detecting regions containing GI tract pathologies. The experiment carried out, and the impact of the results on the advancement of detection models in GI Tract images are also described.
4.1 Dataset Preparation
Initially, the dataset was divided by patient, given that the images were taken from WCE videos and depict frames of the same patient. This division was then designated to prevent data leakage, the influence of extremely similar images of the same patient being present in the same set (training, validation, or test). As a result, only classes with at least three patients were used, since each set must contain at least one different patient with each pathology. In addition, only classes of luminal findings in which the bounding boxes were also available in the dataset were considered. Therefore, based on Table 1, the classes used were Angiectasia, Erosion, Foreign body, Lymphangiectasia, and Ulcer.
As a result, there was an imbalance between the number of pathological images belonging to each patient. So, in order to increase the difference in the number of samples in each set, the patient with the most images was sent to the training set, the second to the validation set, the third to the test set, and so on, in that order, until all the patients in the class were distributed. The number of images with pathology belonging to each of the sets can be seen in Table 2.
In the healthy images used for classification, it was observed that many images were from the same patient. Therefore, efforts were made to diversify the number of normal images obtained from each patient, ensuring a balance with the set of abnormal images. Thus, 200 images from 13 patients were used for the training set, 80 images from 10 patients for validation, and 30 images from 14 patients for testing, totaling 2600 images for training, 800 images for validation, and 420 images for testing.
4.2 Abnormal Image Classification
This section describes the results achieved in the stage of selecting the best-performing CNN architecture. All CNN architectures were trained with 100 epochs and the following hyperparameters: Adam [10] optimizer with a learning rate of 0.001, batch size of 32 and early stopping with patience of 25 epochs. Moreover, the images were normalized and resized to 224 \(\times \) 224. In order to evaluate the models to determine the best performing one, experiments were carried out on the following architectures: VGG16, ResNet50, InceptionV3, Xception, EfficientNetB0, EfficientNetB1, EfficientNetB2, EfficientNetV2B0, EfficientNetV2B1 and EfficientNetV2B2. All the architectures were trained with pre-trained weights from imagenet. Table 3 shows the results achieved with each architecture and the number of parameters in each model.
Although VGG16 has perfect precision, its accuracy, recall and F1-score are low, which indicates that the model makes few positive predictions and is not ideal for applications that require balanced detection. Something similar occurs in EfficientNetB0, as the model has high accuracy but fails to detect part of the true positives.
Xception and EfficientNetV2B2 presented modest results in all metrics, demonstrating that neither model was well-suited to the proposed task. Therefore, ResNet50, InceptionV3 and EfficientNetV2B1 have good accuracy, but their precisions are relatively low, resulting in moderate F1-score values, 78.7% and 70.9%, respectively.
EfficientNetB1 and EfficientNetB2 presented relatively good and balanced precision and recall values, although EfficientNetB2 presented low accuracy compared to the other models. As a result we have 82.7% and 84.5% F1-Score, respectively. For these two architectures, the model does not usually miss a positive instance and also does not make many wrong predictions.
However, EfficientNetV2B0 is the model that presents the best combination of accuracy (87.8%), precision (91.6%), recall (89.1%) and F1-score (90.3%). This indicates that it makes many correct predictions and detects most true positives. In other words, it performs the best correct classification of pathologies among all detections made by the system, thus being the most suitable model to undertake the proposed task of classifying pathological images. Table 4 shows the confusion matrix for the classification with EfficientNetV2B0.
4.3 Detection of Regions with Anomalies
To train the detection models, data augmentation was performed on the images of each class. The operations applied were: Vertical and horizontal flip and blur variations with filter size varying from 1\(\,\times \,\)1 to 7\(\,\times \,\)7 pixels. The results of training the dedicated detection models for each class are shown in Table 5.
The mAP50 returns the average precision of the detections with an IoU threshold of 0.5, while the mAP50-95 returns the results of the mAP with IoU thresholds of 0.5 to 0.95. Finally, the IoU indicates the extent to which the prediction box actually intersected the region with the anomaly. Of particular note were the Angiectasia, Foreign Body and Ulcer classes, which achieved 67.3%, 87.2% and 73.1% IoU, respectively, demonstrating the models’ ability to detect the majority of pathological regions in their respective class.
At the end of training the detection models, the next step is to determine the coordinates of the predicted box with the highest confidence prediction in the images classified as abnormal. Thus, in order to evaluate the model’s prediction in relation to the true region with the pathology, the IoU was used. The results obtained from the predictions per model and the overall evaluation of all the models can be seen in Table 6.
To evaluate overall performance, it should be noted that for normal images, which have passed through the classification filter for detection, a 100% true positive value will be considered if there are no predictions for these images since there are no abnormalities to be found. Otherwise, if there is any kind of prediction in an image without pathology, a 0% true positive value will be considered, as there are no pathologies to be found in such images. In addition, abnormal images that have been classified as normal automatically count as a 0% hit in the method’s overall performance since the detection models won’t have the chance to predict the pathologies present in these images. Some examples of the acquired results can be seen in Fig. 4. The red boxes represent the model’s prediction, while the green boxes show the true anomaly region. In addition, the IoU value of each prediction is highlighted in the top left corner of the images. Although the overall result was not satisfactory, it should be noted that this work represents an initial step in the detection of pathological regions in WCE images, especially with regard to the criteria for dividing up the dataset.
4.4 Discussion
The results shown in Tables 5 and 7 demonstrate the superiority of the dedicated models for each class compared to the single model for all classes in detecting pathologies. For the angiectasia classification, the dedicated model obtained a precision of 94.6% and a mAP50 of 79.3%, in contrast to the single model, which showed a precision of only 21.1% and a mAP50 of 29.9%. Although the recall of the single model is slightly higher, the combination of high precision and mAP50 of the dedicated model indicates a more accurate and reliable identification of angiectasia images. In erosion detection, the dedicated model was also superior, with a precision of 46.6% and a mAP50 of 16.9%, while the single model showed substantially lower values.
For the foreign body class, although both models showed high performances, the dedicated model was still superior in its performance. In the lymphangiectasia and ulcer classes, the dedicated models significantly outperformed the single model in all evaluated metrics. As for the IoU, the dedicated models were superior in detecting regions containing anomalies in all classes. Especially in the erosion and ulcer classes, where the model trained for all classes obtained insignificant results, while the dedicated models could detect more than half of the pathological regions. These results highlight that the approach of training dedicated models for each class not only improves efficiency in detecting specific pathologies, but also reduces the false positive rate and increases confidence in the detections made. Therefore, using dedicated models for pathology detection in WCE is a more effective solution, providing more reliable results than a single model trained for all classes.
In addition, in Table 8 a comparison was made with the binary classification methods of works related to the study carried out in this paper, all using Kvasir-Capsule. Each study contributed different methodologies and made significant advances in the field. However, based on the division criteria adopted by the works that relate to the classification stage carried out in this study, the proposed method still showed good performance compared to state-of-the-art models. In comparison, Muruganantham et al. [17] achieved 94.70% accuracy, Fonseca et al. [5] obtained 97.00% precision, Oukdach et al. [18] achieved F1-Score of 97.00% and Jain et al. [7] achieved 98.00% across all metrics with a hybrid network approach and residual attention, all using the same random division of data.
It is emphasized that the patient splitting criterion adopted in the proposed method is crucial to avoid data leakage between training and testing, ensuring more reliable results when evaluating the model’s performance. This method ensures the integrity of the results by mitigating potential biases introduced by less rigorous data-splitting practices. It is worth noting that during the search for research related to this work, no other work was identified that utilized the Kvasir-Capsule dataset and proposed or evaluated a method to classify abnormalities and detect the region in which the pathology is found. As such, this work aims to propose advances in the area of pathology detection in WCE images of the GI Tract, presenting some baseline results for future work in the area.
5 Conclusion
In conclusion, early diagnosis is essential so that the necessary treatment can be carried out, but the detection of GI Tract pathologies in WCE images remains a challenging task due to the complex nature of the gastrointestinal system and the low quality of the images generated by this exam. The proposed method proved to be a promising tool for detecting GI Tract pathologies present in WCE images. The F1-Score of 90% obtained in the classification stage indicates that the EfficientNetV2B0 model is able to help in the early identification of pathological images, allowing the detection models dedicated to each class to work on the images that present the diseases. Thus, the detection showed the ability to find some of the pathological regions based on the strategy of treating the prediction region as the one with the highest confidence among the trained detection models.
As future work, in order to build on the results obtained in this study, an expansion of the proposed method is intended, with pre-processing methods to improve the models’ ability to identify pathologies. In addition, it is proposed to investigate multi-class classification methods to serve as a step between the first classification and detection, to use the best detection model for the frame analyzed. It is also planned to test the method on other datasets in order to validate the robustness of the proposed method further. Therefore, it is understood that this work can help health specialists in the task of identifying pathologies in WCE images, potentially reducing the workload on doctors and improving patient outcomes through early and accurate diagnosis.
References
Chollet, F.: Xception: deep learning with depthwise separable convolutions. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 1251–1258 (2017)
Chollet, F.: Keras (2015). https://keras.io
Costa, C.L., Lima, D.A., Zorzo Barcelos, C.A., Travençolo, B.A.: Ensemble architectures and efficient fusion techniques for convolutional neural networks: an analysis on resource optimization strategies. In: Brazilian Conference on Intelligent Systems, pp. 107–121. Springer (2023)
Delagah, B., Hassanpour, H., et al.: Feature extraction for polyp detection in wireless capsule endoscopy video frames. J. Healthc. Eng 2023, 6076514 (2023)
Fonseca, F., Nunes, B., Salgado, M., Cunha, A.: Abnormality classification in small datasets of capsule endoscopy images. Procedia Comput. Sci. 196, 469–476 (2022)
He, K., Zhang, X., Ren, S., Sun, J.: Deep residual learning for image recognition. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 770–778 (2016)
Jain, S., Seal, A., Ojha, A.: A hybrid convolutional neural network with meta feature learning for abnormality detection in wireless capsule endoscopy images. arXiv preprint arXiv:2207.09769 (2022)
Jocher, G., Chaurasia, A., Qiu, J.: YOLO by Ultralytics (2023). https://github.com/ultralytics/ultralytics. Accessed 28 Feb 2023
Kaufman, S., Rosset, S., Perlich, C., Stitelman, O.: Leakage in data mining: formulation, detection, and avoidance. ACM Trans. Knowl. Discov. Data 6(4) (2012). https://doi.org/10.1145/2382577.2382579
Kingma, D.P., Ba, J.: Adam: a method for stochastic optimization. arXiv preprint arXiv:1412.6980 (2014)
Li, X., et al.: Generalized focal loss: learning qualified and distributed bounding boxes for dense object detection. Adv. Neural. Inf. Process. Syst. 33, 21002–21012 (2020)
Lima, D.L.S., Pessoa, A.C.P., De Paiva, A.C., da Silva Cunha, A.M.T., Júnior, G.B., De Almeida, J.D.S.: Classification of video capsule endoscopy images using visual transformers. In: 2022 IEEE-EMBS International Conference on Biomedical and Health Informatics (BHI), pp. 1–4. IEEE (2022)
Litjens, G., et al.: A survey on deep learning in medical image analysis. Med. Image Anal. 42, 60–88 (2017)
Mendes, A.C., Pessoa, A.C.P., de Paiva, A.C.: Multi-label classification of pathologies in chest radiograph images using densenet. In: Brazilian Conference on Intelligent Systems, pp. 167–180. Springer (2023)
Milivojevic, V., Milosavljevic, T.: Burden of gastroduodenal diseases from the global perspective. Curr. Treatment Opt. Gastroenterol. 18, 148–157 (2020)
MMYOLO: YOLOv8 by MMYOLO (2023). https://github.com/open-mmlab/mmyolo/tree/main/configs/yolov8. Accessed 15 Mar 2024
Muruganantham, P., Balakrishnan, S.M.: Attention aware deep learning model for wireless capsule endoscopy lesion classification and localization. J. Med. Biol. Eng. 42(2), 157–168 (2022)
Oukdach, Y., Kerkaou, Z., El Ansari, M., Koutti, L., Fouad El Ouafdi, A., De Lange, T.: Vitca-NDet: a framework for disease detection in video capsule endoscopy images using a vision transformer and convolutional neural network with a specific attention mechanism. Multimedia Tools Appl. 1–20 (2024)
Qureshi, R., et al.: A comprehensive systematic review of yolo for medical object detection (2018 to 2023). Authorea Preprints (2023)
Simonyan, K., Zisserman, A.: Very deep convolutional networks for large-scale image recognition. arXiv preprint arXiv:1409.1556 (2014)
Smedsrud, P.H., et al.: Kvasir-capsule, a video capsule endoscopy dataset. Sci. Data 8(1), 142 (2021)
Srivastava, A., Tomar, N.K., Bagci, U., Jha, D.: Video capsule endoscopy classification using focal modulation guided convolutional neural network. In: 2022 IEEE 35th International Symposium on Computer-Based Medical Systems (CBMS), pp. 323–328. IEEE (2022)
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., Wojna, Z.: Rethinking the inception architecture for computer vision. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 2818–2826 (2016)
Tan, M., Le, Q.: EfficientNet: rethinking model scaling for convolutional neural networks. In: International Conference on Machine Learning, pp. 6105–6114. PMLR (2019)
Tan, M., Le, Q.: Efficientnetv2: smaller models and faster training. In: International Conference on Machine Learning, pp. 10096–10106. PMLR (2021)
Valentim, N.A., Dorça, F.A., Asnis, V.P., Elias, N.C.: The artificial intelligence as a technological resource in the application of tasks for the development of joint attention in children with autism. In: Brazilian Conference on Intelligent Systems, pp. 306–320. Springer (2023)
World Health Organization: The Global Cancer Observatory (2024). https://gco.iarc.fr/en. Accessed 15 Mar 2024
Xu, Z., et al.: Upper gastrointestinal anatomy detection with multi-task convolutional neural networks. Healthcare Technol. Lett. 6(6), 176–180 (2019)
Acknowledgments
The authors acknowledge the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES), Brazil - Finance Code 001, Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Brazil, and Fundação de Amparo à Pesquisa Desenvolvimento Científico e Tecnológico do Maranhão (FAPEMA) (Brazil), Empresa Brasileira de Serviços Hospitalares (Ebserh) Brazil (Grant number 409593/2021-4) for the financial support.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2025 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Silva, A.L. et al. (2025). Detection of Pathological Regions of the Gastrointestinal Tract in Capsule Images Using EfficientNetV2 and YOLOv8. In: Paes, A., Verri, F.A.N. (eds) Intelligent Systems. BRACIS 2024. Lecture Notes in Computer Science(), vol 15412. Springer, Cham. https://doi.org/10.1007/978-3-031-79029-4_23
Download citation
DOI: https://doi.org/10.1007/978-3-031-79029-4_23
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-79028-7
Online ISBN: 978-3-031-79029-4
eBook Packages: Computer ScienceComputer Science (R0)