Abstract
Large Language Models (LLMs) have been widely employed in various text processing tasks. In computer vision, these models have found application in generating captions and text from natural images, as well as in Visual Question Answering (VQA) systems. In the field of medical imaging, there are studies based on text generation proposing automated diagnoses of X-rays, magnetic resonance imaging scans, computed tomography scans, and other modalities. Few initiatives seek to apply and harness the potential of LLMs in medical text generation; they use models with tens of billions of parameters and are thus computationally expensive. This work addresses this gap by evaluating the use of frozen pre-trained models (CXAS U-Net and BioGPT) for chest X-ray report generation. We adapt the BLIP-2 modular architecture where only a cross-modal alignment module must be trained in order to generate text from images. We were able to achieve competitive scores over Clinical Efficacy (CE) metrics compared to some state-of-the-art (SOTA) methods, while obtaining lower scores for Natural Language Generation (NLG) metrics. Our findings suggest that NLG metrics may not serve as suitable proxies for evaluating models in the chest X-ray generation task.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Chest X-rays are a fundamental diagnostic tool in healthcare, containing valuable information that can assist healthcare professionals in diagnosing various pulmonary, cardiac or traumatic conditions [1]. They provide a snapshot of a patient’s internal chest structure, revealing critical details that can guide medical practitioners in their decision-making processes.
However, interpreting and extracting the findings from chest X-rays is a complex, repetitive and often time-consuming task. The analysis of these images traditionally relies on the expertise of radiologists, who interpret the visual information contained within the X-rays and then transcribe their findings into textual reports. This manual process is not only labor-intensive but is also prone to variations in interpretation and reporting. It is in this context that innovative solutions are needed to enhance the efficiency, accuracy, and accessibility of chest X-ray analysis [12].
In recent years, the advancements in artificial intelligence (AI) and natural language processing (NLP) are revolutionizing the way humans interact with the machines [20, 31]. Some Large Language Models (LLMs) applications, such as ChatGPT, are showing off their exceptional text generation capabilities, being able to output texts with impressive syntax and semantic quality. They also show a good performance while performing tasks they were not previously trained for, for example, one can prompt them to generate a clinical diagnostic, or write a code to solve a given problem.
Several works are harnessing the developments in AI and NLP to automate and streamline the process of transforming visual information from chest X-rays into informative textual descriptions, however, only a few have capitalized on the potential of LLMs. These existing methods predominantly rely on resource-intensive models that present challenges in accessibility for those with limited computational resources.
Therefore, to leverage the capabilities of LLMs while minimizing the resources required for training and deploying a chest X-ray report generation system, this study proposes adapting the BLIP-2 modular architecture [16] to utilize frozen in-domain pre-trained models for both vision encoding and text generation. We employ BioGPT [18] as our text generation model, which is based on GPT-2 with 354M parameters, offering significantly lower resource requirements compared to models used in other studies [6, 30, 36] and widely-used LLMs currently used deep learning area [20, 31].
By adopting this modular architecture, where the image encoder and text decoder weights are frozen, only a vision-text alignment module needs to be trained, reducing training complexity and computational costs compared to having to train a large LLM.
By harnessing the power of LLMs while seeking computational efficiency, this research contributes to the advancement of AI-powered medical image analysis. This advancement has the potential to reduce radiologists’ workload by serving as a decision support, increasing throughput, minimizing diagnosis delays, and allowing them to focus on complex analyses where their expertise is most valuable [11].
2 Background
The literature review focuses on three areas: Image Captioning, Medical Captioning, and Language Models. In the first two areas, we make an comprehensive exploration focusing on different approaches, architectures, benchmarks and applicable metrics for our task. Lastly, we investigate some Language Models, aiming to identify their sizes and number of parameters.
2.1 Image Captioning
The image captioning task targets the automatic generation of description of natural images. In the literature, we find many methods designed to achieve this goal [16, 17, 23, 33, 37, 39]. These works vary based on: their selection of vision encoder, language model, and their approach to cross-domain alignment, which involves aligning features between the image and text domains.
Vinyals et al. [33] and Rennie et al. [23] used CNN models to extract the features from images and LSTM network to regressively generate the desired text. The CNN features provide a simple and compact representation of an image, but they can hinder further fine-grained description due to this compression. The LSTM network, due to its sequential nature, can be very slow to output the text.
Zhou et al. [37] and Li et al. [17] leveraged the potential of the Transformer [32] along region image features extracted by Fast-RCNN [9], an widely used CNN for feature extraction. Zhou et al. [37], for example, used a shared multi-layer Transformer responsible for both vision encoding and text decoding steps, hence this unique module is responsible for aligning the image features and generating the caption.
Very recent works [16, 39] take advantage of the few shot potential of LLMs [3]. Zhu et al. [39] introduce MiniGPT-4, which combines a Vision Transformer image encoder with an open-source LLM based on Llama [31], aligning both image and text domain using solely a single projection layer. The authors named it MiniGPT-4 due to its similar capability on description generation compared to GPT-4.
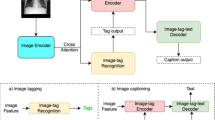
Built on pre-trained models, Li et al. [16] introduced the BLIP-2 framework, which requires training only an cross-modal alignment module called Q-Former to bridge the gap between image and text representations. This approach leverages pre-trained image encoders and large language models (LLMs) by freezing their weights (i.e., not training them).
The core component, Q-Former, is a BERT-based module responsible for aligning the features extracted from images and text. By training only the Q-Former, BLIP-2 reduces training complexity and computational costs.
2.2 Medical Captioning
Medical captioning involves the automatic generation of descriptive and informative captions for medical images, such as X-rays, MRI scans, CT scans, and more. The goal is to provide accurate textual descriptions that convey relevant medical information, possibly helping medical professionals make faster and more accurate diagnoses, facilitating research and training, and ultimately improving patient care. Considerable progress has been made in this field, with numerous proposed architectures and approaches [4,5,6, 26, 30, 35, 36].
Some works, particularly those focused on chest X-ray captioning, employ CNNs as image encoders and Transformers as text decoders [4, 5]. Chen et al. [4] introduced the use of a novel relational memory module alongside the Transformer to enhance the caption generation, while Chen et al. [5] focused on the alignment of image and text domain, proposing the use of a cross-modal memory network to facilitate the interactions across modalities (image and text). On the other hand, a recent study [35] adopts a vision transformer as its image encoder and introduces Expert Tokens, which are designed to interact with the extracted images patches and with one another. As a result, each token can focus on a different part of the image.
Recently, other papers have started to explore the potential of LLMs for medical caption generation [6, 30, 36]. Yang et al. [36] adopted the BLIP-2 architecture, although they do not follow the original training process, instead, they directly use a large vision encoder and a LLM decoder, and fine-tune the Q-Former and the LLM to the ImageClef task, which involves caption prediction for general X-ray images. They compared their performance solely based on the task competition ranking.
Thawkar et al. [30] used the Vision Transformer from MedCLIP [34] as the vision encoder, applied a simple linear transformation layer to align features, and employed an LLM based on LLaMA [31] as the text decoder. This study does not compare their approach with others. Danu et al. [6] took a different approach by first multi-classifying diseases on chest X-ray images with bounding boxes, and then using these abnormality features as input for the LLM.
Med-PaLM [26, 27] is a large-scale generalist biomedical AI system, capable of interpreting various biomedical data modalities, including tasks like chest X-ray report generation and medical visual question answering. It employs a ViT-based vision encoder and PaLM as the LLM decoder.
Building on prior work, Nicolson et al. [19] investigated the impact of combining different pre-trained models for chest X-ray report generation. They experimented with various combinations of vision encoders and text decoders, both from the medical domain (in-domain) and general domains. Interestingly, their findings revealed that a combination of an out-of-domain vision encoder, CvT-21, and a general-domain text decoder, DistilGPT-2, achieved superior performance on chest X-ray report generation benchmarks after fine-tuning. This finding suggests that leveraging out-of-domain models, when carefully chosen and fine-tuned, can lead to state-of-the-art (SOTA) results.
2.3 Language Models
In recent years, significant advancements have been made in the development of language models. A significant milestone was the introduction of BERT [8] (Bidirectional Encoder Representations from Transformers), which has 110 million parameters in its base version and 340 million in its large version. BERT’s bidirectional training approach on masked language modeling tasks enabled it to achieve state-of-the-art results on a variety of NLP benchmarks.
The transformer-based architectures have been further scaled to develop models with even larger parameter sizes. For instance, OpenAI’s GPT-2 [21], with 1.5 billion parameters on its larger model, demonstrated the potential of unsupervised language models to generate coherent and contextually relevant text. GPT-2’s architecture paved the way for its successor, GPT-3 [3], which boasts an unprecedented 175 billion parameters, showcasing remarkable capabilities in few-shot learning and diverse NLP tasks without requiring task-specific fine-tuning.
Despite their impressive performance, these large-scale models necessitate substantial computational resources for training and inference, limiting their accessibility. This challenge has driven the development of more resource-efficient models. DistilBERT [24], with 66 million parameters, and ALBERT [14], with 12 million parameters for its base version, offer more compact alternatives that maintain competitive performance levels.
In the biomedical domain, specialized language models have been developed to address the unique challenges posed by medical texts. BioBERT [15], based on BERT with approximately 110 million parameters, and ClinicalBERT [2], also based on BERT, have been fine-tuned on large corpora of biomedical literature and clinical notes, resulting in improved performance on domain-specific tasks.
Following the LLMs line, BioGPT [18], a model derived from GPT-2 with 354 million parameters, specifically tailored for biomedical text generation, provides a balanced approach, offering the extensive capabilities of larger models while maintaining lower resource requirements. This efficiency makes BioGPT a practical choice for potentially generating high-quality biomedical text, ensuring accessibility for a broader range of researchers.
Wrapping up, we observe that many works in medical captioning, specifically chest X-ray captioning, are focused on enhancing performance metrics. Furthermore, those concentrating on harnessing the power of LLMs do not appear to prioritize the search for reduced-size architectures that would enable simple computers with limited computational resources to run these models.
3 Materials
This work adopts the MIMIC-CXR dataset [13], a widely-used dataset for generating captions for chest X-rays. It is currently the largest publicFootnote 1 chest x-ray dataset, containing 377,100 radiograms and 227,835 free-text reports. The dataset has an official data split allowing fair comparison of our models against literature methods and different techniques.
Most related works also utilize the IU Xray dataset [7]. However, in this study, we have chosen to use only the MIMIC-CXR due to its larger size. In the future, we plan to include additional datasets to assess generalization.
Since MIMIC-CXR includes sample reports with multiple views, such as frontal and lateral X-rays, we employed the following strategy for inference:
-
 For reports with two images: We randomly selected two images for analysis.
For reports with two images: We randomly selected two images for analysis. -
 For reports with only one image: We duplicated the single image to ensure the pipeline could process it consistently.
For reports with only one image: We duplicated the single image to ensure the pipeline could process it consistently.
Following this selection process, all images were resized to a standard dimension of 512\(\times \)512 pixels. Additionally, normalization was applied using the mean and variance values specific to the chosen vision encoder.
4 Method
This section presents the proposed model architecture, the training stages, and the metrics used during the evaluation process.
Model Architecture. This works leverages the architecture and training pipeline presented by BLIP-2 [16], a captioning model for natural images that utilizes frozen pre-trained models.
The key advantage of this architecture is its modularity. This allows for the integration of different vision encoders and text decoders, with only the alignment module (Q-Former) requiring specific training. The source code was based and adapted from BLIP-2 implementation on LAVIS repositoryFootnote 2.
Image Encoder Stage. The architecture, as illustrated in Fig. 1, starts with the image encoder, responsible for extracting pertinent features from chest X-ray images.
This work adopted a U-Net [25] formerly pre-trained on a chest x-ray segmentation dataset, therefore we can leverage its prior in-domain knowledge. This choice offers a significant advantage, as U-Net has nearly 15 times fewer parameters compared to the Vision Transformer (ViT-g/14) employed in the original BLIP-2 work. At this point, we utilized the feature map output of the first upsampling layer in the network, flattened the height and width dimensions, and then switched channels dimension with this flattened spatial dimension. This allows having features that are ready to be processed by the next neural network.
Regarding the image feature extraction, the study utilizes two images for every sample as aforementioned, hence we followed other studies [4, 35], sequentially extracting features from both images using the selected image encoder, and averaging those features.
Q-Former Alignment Module. The cornerstone of the architecture, The Q-Former is responsible for aligning extracted images features with the text decoder input.
As the Q-Former is a BERT-based model, we leveraged available BioClinicalBERT [2] pre-trained weights, which was trained on PubMed abstracts and also on MIMIC III notes. Initializing language models with in-domain pre-trained weights, instead of using the standard BERT initialization, potentially boosts the capacity of feature alignment of the model; behavior seen in other medical tasks [15]. We used the same configuration from the base source code.
Text Decoder Module. The text decoder is the module responsible for generating the chest X-ray descriptions given the features aligned by the Q-Former.
Similar to BLIP-2 architecture, we adopted a Large Language Model as our text decoder, due to its text generation capability. Generally, an LLM receives text prompts on its input which work as an start point, from where the model generates the text output. However, in this work, as the LLM inputs come from the Q-Former aligned features, these inputs can be considered as soft prompts, i.e., they are not explicit text embeddings, but a feature vector that must help and guide the LLM decoder on its text generation task.
This work utilizes BioGPT [18], an LLM based on GPT-2 [21] architecture, since it was specifically pre-trained on biomedical domain using PubMed abstracts. It is a lightweight model compared to the LLM from the original BLIP-2, containing 345 million parameters instead of 2.7 billion.
4.1 Training Stages
The training process is divided into two distinct stages, each serving a specific purpose in enhancing the capabilities of a portion of our architecture, as illustrated in Fig. 1.
Representation Learning. In this initial training stage, the Q-Former is trained in conjunction with a frozen pre-trained image encoder. The primary objective is to enable the Q-Former to serve as a bridge between image and text domains. This step prepares the module to align, understand, and also, generate meaningful captions, enhancing the performance of the architecture as a whole. During this phase, the Q-Former outputs are optimized using three distinct methods (cost functions), as outlined in the BLIP-2 work [16].
We adopted AdamW optimizer with weight decay of 0.05. The learning rate followed a cosine decay schedule during 10 epochs, starting at 1e-4 and ending at 1e-5. A warm-up period of one epoch was employed with learning rate set to 1e-6. Also, the best model checkpoint was selected based on the lowest loss overall.
Generative Learning. In the second training stage, our whole architecture is assembled. The previously trained Q-Former module is integrated with the frozen pre-trained BioGPT. Subsequently, the Q-Former is fine-tuned by optimizing the outputs generated by the LLM decoder. During this fine-tuning process, the Q-Former acts as a guide, influencing BioGPT’s text generation based on the input chest X-ray images.
We adopted AdamW optimizer with weight decay of 0.05. The learning rate followed a cosine decay schedule during 10 epochs, starting at 1e-5 and ending at 1e-6. A warm-up period of two epochs was employed with learning rate set to 1e-8. At this point, we observed better results by unfreezing the BioGPT and freezing all the other modules. Same as before, the final model checkpoint was selected based on the lowest loss value. All training stages were performed on AWS Cloud using a ml.g5.xlarge instance containing a 24Gb GPU and 16Gb memory.
4.2 Metrics and Evaluation
Following Chen et al. [4, 35, 38], our evaluation aimed to assess both the quality of the generated text descriptions and their potential clinical usefulness.
We adopted two approaches, leveraging established Natural Language Generation (NLG) metrics (BLEU, METEOR and ROUGE-L) and additionally, we incorporated clinical efficacy (CE), where we employed the CheXbert system [28] to automatically label the generated reports within 14 categories.
By comparing these labels with the ground truth diagnoses, we calculated precision, recall, and F1-score, providing insights into the model’s ability to generate clinically accurate reports. This work presents both micro and macro averages of precision, recall and F1-score [29].
5 Experimental Results
This section presents the findings of our evaluation on the effectiveness of the proposed method for generating clinical text reports. We assessed performance using a combination of Natural Language Generation (NLG) metrics and clinical efficacy (CE) metrics over the MIMIC-CXR dataset.
Our architecture consists of approximately 450 million parameters. In comparison with other techniques that also adopt LLM in the decoder stage, XRayGPT [30] has around 7 billion parameters solely in its large language model (LLM). Similarly, Yang et al. [36] employs the ChatGLM-6B LLM model, which comprises approximately 6 billion parameters. Med-PaLM [26], on the other hand, incorporates 540 billion parameters in its PaLM LLM. Unfortunately, these studies do not evaluate their models using commonly used datasets, thus our comparison is limited to the number of parameters.
In order to quantitatively compare our approach with other studies in the area, we evaluated the clinical efficacy of the reports using CheXbert labels. Precision, recall, and F1-score were calculated for each CheXbert finding. Table 1 summarizes the overall CE performance compared to existing methods.
Our method achieved the highest precision (0.504) among the evaluated methods, indicating a low rate of false positive findings. However, the recall score (0.346) suggests it may overlook some relevant findings compared to CvT21-2DistilGPT2 [19].
Table 2 presents a comparison of NLG metrics between other techniques and our model. Our model has low scores if compared to other techniques. This could be explained due to the fact the those NLG metrics were developed to mainly evaluate machine translation, which is clearly not the task here. Since our core model is a pre-trained GPT-2 model, our predictions may not align as closely with the ground truth. Consequently, this misalignment could account for the observed low scores in our proposed model’s metrics.
A further analysis revealed an interesting nuance between clinical efficacy (CE) metrics and Natural Language Generation (NLG) metrics in the context of chest X-ray report generation. Specifically, within a subset of high-performing predictions based on CE metrics—where predictions effectively conveyed observations from the X-rays—the corresponding NLG metrics displayed lower scores (see Table 3). This observation suggests that NLG metrics alone may not be the most reliable indicator of model quality for this task.
Even predictions which are clinically accurate by CE metrics might not receive high NLG scores. This highlights the limitations of NLG metrics in capturing the full spectrum of factors that contribute to good clinical reports, such as factual accuracy, nuanced phrasing, and adherence to reporting conventions.
Finally, as shown in Table 4, the model performs well on identifying findings such as Support Devices (0.683 F1) and Cardiomegaly (0.521 F1). However, it struggles with less prevalent findings like Atelectasis (0.330 F1) and Lung Opacity (0.229 F1). This may be due to limitations in the training data or the model’s capacity to capture subtle variations in these specific findings.
These results demonstrate the potential of our proposed method for generating clinically relevant and accurate text reports. The high precision ensures a low false positive rate, while further investigation is needed to improve recall, particularly for less frequent findings.
6 Conclusions
In conclusion, our work presented an approach for generating clinical text reports from chest X-ray images. The selection of relative small-sized models, and the modular characteristic of BLIP-2 has made the training process less-hardware intensive, and we hope it foster future investigations of this type of architecture.
Our method achieved the highest precision among the evaluated techniques. However, our evaluations also reveals challenges to achieve a high recall, particularly for less prevalent observations. Hence, the model needs further investigations to improve model’s capacity.
Furthermore, our analysis of the NLG metrics highlights the limitations of using machine translation metrics for our specific task, which do not capture the actual quality of the report. Future efforts should focus on developing custom evaluation methods that align with our specific goals.
Overall, our results demonstrate the potential of using in-domain, pre-trained, reduced-size models to generate clinically relevant and accurate text reports. However, ongoing research and development efforts are still needed to improve model’s recall, to refine to overall’s capability of the model. With this improvements, this approach could become a valuable tool, contributing to more efficient and accurate diagnosis.
As future directions, we aim to assess this framework with other datasets, such as BRAX [22] and IU XRay [7], to validate model’s generalization.
Notes
- 1.
MIMIC-CXR webpage: https://physionet.org/.
- 2.
LAVIS repository: https://github.com/salesforce/LAVIS.
References
Al-Dasuqi, K., Johnson, M.H., Cavallo, J.J.: Use of artificial intelligence in emergency radiology: an overview of current applications, challenges, and opportunities. Clin. Imaging 89, 61–67 (2022)
Alsentzer, E., Murphy, J.R., Boag, W., Weng, W.-H., Jin, D., Naumann, T., McDermott, M.: Publicly available clinical BERT embeddings. arXiv preprint arXiv:1904.03323 (2019)
Brown, T., et al.: Language models are few-shot learners. Adv. Neural. Inf. Process. Syst. 33, 1877–1901 (2020)
Chen, Z., Song, Y., Chang, T.-H., Wan, X.: Generating radiology reports via memory-driven transformer. arXiv preprint arXiv:2010.16056, 2020
Chen, Z., Shen, Y., Song, Y., Wan, X.: Cross-modal memory networks for radiology report generation. arXiv preprint arXiv:2204.13258, 2022
Danu, M.D., et al.: Generation of Radiology Findings in Chest X-Ray by Leveraging Collaborative Knowledge. arXiv preprint arXiv:2306.10448 (2023)
Demner-Fushman, D., et al.: Preparing a collection of radiology examinations for distribution and retrieval. J. Am. Med. Inf. Assoc. 23(2), 304–310 (2016)
Devlin, J., Chang, M.-W., Lee, K., Toutanova, K.: BERT: pre-training of deep bidirectional transformers for language understanding. arXiv preprint arXiv:1810.04805 (2018)
Girshick, R.: Fast R-CNN. In: IEEE International Conference on Computer Vision, pp. 1440–1448 (2015)
Gu, T., Liu, D., Li, Z., Cai, W.: Complex organ mask guided radiology report generation. In: IEEE/CVF Winter Conference on Applications of Computer Vision, pp. 7995–8004 (2024)
Irmici, G., et al.: Chest x-ray in emergency radiology: what artificial intelligence applications are available? Diagnostics 13(2), 216 (2023)
Jalal, S., Parker, W., Ferguson, D., Nicolaou, S.: Exploring the role of artificial intelligence in an emergency and trauma radiology department. Can. Assoc. Radiol. J. 72(1), 167–174 (2021)
Johnson, A.E., et al.: MIMIC-CXR-JPG, a large publicly available database of labeled chest radiographs. arXiv preprint arXiv:1901.07042 (2019)
Lan, Z., et al.: Albert: A lite bert for self-supervised learning of language representations. arXiv preprint arXiv:1909.11942 (2019)
Lee, J., et al.: BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36(4), 1234–1240 (2020)
Li, J., Li, D., Savarese, S., Hoi, S.: BLIP-2: bootstrapping language-image pre-training with frozen image encoders and large language models. arXiv preprint arXiv:2301.12597 (2023)
Li, X., et al.: Oscar: object-semantics aligned pre-training for vision-language tasks. In: Vedaldi, A., Bischof, H., Brox, T., Frahm, J.-M. (eds.) ECCV 2020. LNCS, vol. 12375, pp. 121–137. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-58577-8_8
Luo, R., et al.: BioGPT: generative pre-trained transformer for biomedical text generation and mining. Brief. Bioinform. 23(6), 409–421 (2022)
Nicolson, A., Dowling, J., Koopman, B.: Improving chest X-ray report generation by leveraging warm starting. Artif. Intell. Med. 144, 102633 (2023)
OpenAI. GPT-4 Technical Report (2023)
Radford, A., Wu, J., Child, R., Luan, D., Amodei, D., Sutskever, I., et al.: Language models are unsupervised multitask learners. OpenAI Blog 1(8), 9 (2019)
Reis, E.P., et al.: Brax, brazilian labeled chest x-ray dataset. Sci. Data 9(1), 487 (2022)
Rennie, S.J., Marcheret, E., Mroueh, Y., Ross, J., Goel, V.: Self-critical sequence training for image captioning. In: IEEE Conference on Computer Vision and Pattern Recognition (2017)
Sanh, V., Debut, L., Chaumond, J., Wolf, T.: DistilBERT, a distilled version of BERT: smaller, faster, cheaper and lighter. arXiv preprint arXiv:1910.01108 (2019)
Seibold, C., et al.: Accurate fine-grained segmentation of human anatomy in radiographs via volumetric pseudo-labeling. arXiv preprint arXiv:2306.03934 (2023)
Singhal, K., et al.: Large language models encode clinical knowledge. arXiv preprint arXiv:2212.13138 (2022)
Singhal, K., et al.: Towards expert-level medical question answering with large language models. arXiv preprint arXiv:2305.09617 (2023)
Smit, A., et al.: Chexbert: combining automatic labelers and expert annotations for accurate radiology report labeling using bert. arXiv preprint arXiv:2004.09167 (2020)
Sorower, M.S.: A literature survey on algorithms for multi-label learning. Oregon State University, Corvallis 18(1), 25 (2010)
Thawkar, O., et al.: XRayGPT: chest radiographs summarization using medical vision-language models. arXiv preprint arXiv:2306.07971 (2023)
Touvron, H., et al.: Llama 2: open foundation and fine-tuned chat models. arXiv preprint arXiv:2307.09288 (2023)
Vaswani, A., et al.: Attention is all you need. Adv. Neural Inf. Process. Syst. 30 (2017)
Vinyals, O., Toshev, A., Bengio, S., Erhan, D.: Show and tell: a neural image caption generator. In: IEEE Conference on Computer Vision and Pattern Recognition, pp. 3156–3164 (2015)
Wang, Z., Wu, Z., Agarwal, D., Sun, J.: MedCLIP: contrastive learning from unpaired medical images and text. arXiv preprint arXiv:2210.10163 (2022)
Wang, Z., Liu, L., Wang, L., Zhou, L.: METransformer: radiology report generation by transformer with multiple learnable expert tokens. In: IEEE/CVF Conference on Computer Vision and Pattern Recognition, pp. 11558–11567 (2023)
Yang, B., Raza, A., Zou, Y., Zhang, T.: Customizing general-purpose foundation models for medical report generation. arXiv preprint arXiv:2306.05642 (2023)
Zhou, L., Palangi, H., Zhang, L., Hu, H., Corso, J., Gao, J.: Unified vision-language pre-training for image captioning and VQA. In: AAAI Conference on Artificial Intelligence, vol. 34, pp. 13041–13049 (2020)
Zhou, Z., Shi, M., Wei, M., Alabi, O., Yue, Z., Vercauteren, T.: Large model driven radiology report generation with clinical quality reinforcement learning. arXiv preprint arXiv:2403.06728 (2024)
Zhu, D., Chen, J., Shen, X., Li, X., Elhoseiny, M.: MiniGPT-4: enhancing vision-language understanding with advanced large language models. arXiv preprint arXiv:2304.10592 (2023)
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2025 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Viana Vargas, T., Pedrini, H., Santanchè, A. (2025). LLM-Driven Chest X-Ray Report Generation With a Modular, Reduced-Size Architecture. In: Paes, A., Verri, F.A.N. (eds) Intelligent Systems. BRACIS 2024. Lecture Notes in Computer Science(), vol 15413. Springer, Cham. https://doi.org/10.1007/978-3-031-79032-4_14
Download citation
DOI: https://doi.org/10.1007/978-3-031-79032-4_14
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-79031-7
Online ISBN: 978-3-031-79032-4
eBook Packages: Computer ScienceComputer Science (R0)



 For reports with two images: We randomly selected two images for analysis.
For reports with two images: We randomly selected two images for analysis. For reports with only one image: We duplicated the single image to ensure the pipeline could process it consistently.
For reports with only one image: We duplicated the single image to ensure the pipeline could process it consistently.