Abstract
Breast cancer classification in ultrasound images is a challenging and arduous task, primarily due to the quality of the images and the complexity of the lesions encountered. Early diagnosis of this pathology is known to increase patient survival chances. Therefore, computational methods have been developed to assist specialist physicians in this critical task. This study introduces AnisotropicBreast-ViT, a method that integrates anisotropic filtering, balanced data augmentation, and Vision Transformer to aid in the classification of breast ultrasound images. The proposed approach achieves promising results with an accuracy of 98.82%, specificity of 98.62%, sensitivity of 99.25%, precision of 97.08%, F1-score of 98.16%, and AUC-ROC of 0.989, surpassing current benchmarks in the field. These findings suggest that AnisotropicBreast-ViT has the potential to significantly improve breast cancer diagnosis, demonstrating its effectiveness and robustness in clinical applications.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Breast cancer is one of the leading causes of mortality among women worldwide. According to the World Health Organization (WHO), approximately 15% of annual deaths are attributed to this disease [3]. Early detection is crucial for effective treatment, thereby contributing to the reduction of mortality rates. Ultrasonography (US) is one of the safe techniques used for the detection and diagnosis of breast cancer, as it is a non-invasive method, does not involve radiation, and provides high-resolution images in real-time [5].
However, US image interpretation requires the expertise of highly trained radiologists. Even the most experienced radiologists can exhibit variations in their tumor diagnoses, leading to inconsistencies [27]. To address these challenges, computer-aided detection and diagnosis systems (CAD/CADx) have emerged. These systems have the potential to automate the analysis of US images, providing support to healthcare professionals in the identification and classification of breast lesions [24].
Furthermore, deep learning methods such as convolutional neural networks (CNNs) have proven to be highly effective in various computer vision applications in the medical field [11, 12, 16, 17, 23]. These models are capable of identifying patterns directly from images, eliminating the need for manual feature extraction [8]. However, it is important to note that US images are complex and often require appropriate preprocessing to highlight suspicious structures, thus ensuring accurate results.
Considering this issue, the main goal of this work is to develop and evaluate an automated method for breast lesion classification in ultrasound images using image preprocessing techniques and deep learning. Also, the use of anisotropic and mean filters to improve the classification pipeline of breast cancer in medical images. The primary contributions of the proposed method include:
-
Development of an automated classification method that uses image enhancement techniques to highlight intrinsic features (anisotropic and mean filters) of lesions in ultrasound images;
-
Creation of a balanced data augmentation method for the effective training of a Vision Transformer;
-
Robust method composed of techniques that optimize the results of the Vision Transformer, improving classification reliability.
2 Related Works
Breast cancer classification in ultrasound images is widely studied in the literature. In this section, we present the main related works to the proposed method found from the year 2020 onwards. It is worth noting that the search was also conducted considering the variability of existing techniques to identify gaps among them.
In [2], the authors highlight the challenge of training deep learning models due to the scarcity of training samples in breast ultrasound (US) images. To address this problem, the authors proposed an approach called noise filter network (NF-Net). This approach demonstrated an accuracy of 73%, sensitivity of 80%, and F1-score of 74%. The results suggest that NF-Net is a promising approach for breast lesion classification, showing the potential to improve accuracy in scenarios with limited data.
In [24], breast lesion classification in ultrasound (US) images was performed using image fusion and various CNN architectures, such as VGGNet, ResNet, and DenseNet. The authors achieved notable results, with an accuracy of 94.62%, sensitivity of 92.31%, specificity of 95.60%, precision of 90%, F1-score of 91.14%, and an area under the curve (AUC) of 0.9711. These results, obtained on the BUSI dataset, indicate the effectiveness of image fusion techniques and various CNNs in detecting breast lesions.
In the study conducted by [25], a Radiomics-based technique combined with a semi-supervised adversarial neural network model for data augmentation was used. The dataset, of private origin, contained 1,447 US images, with 767 benign and 680 malignant. The best results achieved were an accuracy of 90.41%, sensitivity of 87.94%, and specificity of 85.86%. These results highlight the potential of using Radiomics and semi-supervised learning to improve the classification of breast lesions in the US.
The authors of [31] investigated bilateral filters with various CNN architectures to determine the best training model. The study results showed an accuracy of 95.48%, sensitivity of 93.92%, specificity of 98.33%, precision of 98.11%, F1-score of 95.71%, and AUC of 0.9883. These results indicate that preprocessing with bilateral filtering, combined with optimized CNN architectures, can significantly improve the accuracy of breast lesion classification in the US.
In the study by [15], a hybrid method was presented that uses three CNNs for feature extraction. Then, the Minimum Redundancy Maximum Relevance algorithm is applied to select the most relevant features, and a Support Vector Machine (SVM) is used to train the model. This method achieved an accuracy of 95.6%, highlighting the effectiveness of combining feature extraction and selection techniques in improving the performance of classification models.
In turn, [28] proposed an architecture based on InceptionV3 for breast lesion classification. To overcome the lack of data, the authors used five combined datasets. The resulting model achieved an accuracy of 81%, sensitivity of 77%, F1-score of 80%, and AUC of 0.81. These results suggest that using multiple datasets can help mitigate issues related to data scarcity and improve the robustness of deep learning models.
Finally, [21] proposed using three combined ResNet architectures for feature extraction. After extraction, three different techniques were applied to select the most relevant features, and training was performed using an SVM. Using a dataset with 1,236 images, the best results achieved were an accuracy of 84.9%. These results indicate that combining multiple CNN architectures and feature selection techniques can improve the performance of breast lesion classification models.
The related works present various approaches for classifying breast lesions in ultrasound images, each with its advancements. Techniques range from methods involving Radiomics and semi-supervised learning to methods using CNNs and image fusion.
Despite significant advancements, each study faces specific challenges, such as the scarcity of training data, variations in image quality, and the need for intensive preprocessing. The method proposed in this work aims to fill a crucial gap by employing advanced preprocessing techniques and deep learning to automate and enhance breast lesion classification in ultrasound images. By integrating image enhancement techniques and a balanced data augmentation method for training a Vision Transformer, our method aims to achieve higher classification accuracy and robustness. Additionally, the proposed approach seeks to simplify the process for healthcare professionals, providing faster and more accurate diagnosis, which is crucial for early detection and breast cancer treatment.
3 Materials and Proposed Method
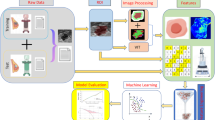
In this section, we describe the materials and the proposed method. It consists of five steps, as highlighted in Fig. 1. Subsequently, we detail each step.
3.1 Materials (BUSI Dataset)
The dataset used for this study was the Breast Ultrasound Images Dataset (BUSI). The BUSI was collected in 2018 and includes women aged between 25 and 75 years. The dataset has 780 images with an average size of \(500\times 500\) pixels in PNG format. Also, a lesion marking file accompanies each image and has three classes: normal, benign, and malignant [1]. Only images from the benign (210 images) and malignant (454 images) classes were selected in this work. This choice is because the fact that a biopsy is only necessary when there is doubt in the diagnosis between benign and malignant. Therefore, normal images were not included in the dataset, focusing on the problem of diagnosing benign and malignant lesions, which is the main objective of this study. Figure 2 shows examples of each class.
In the next step, we will describe the extraction of the Region of Interest (ROI), which was used to enhance the model by eliminating unnecessary information from the images.
3.2 ROI Cropping
As described earlier, the images contain markings made by specialists. It is also noticeable, as illustrated in Fig. 2, that there is a significant amount of information in the image that is not essential for classification and may even confuse the classifier. Many studies show improved results using ROI extraction [6, 7, 9, 11]. Therefore, after conducting detailed experiments, we chose ROI crop. In practice, this involves selecting the largest bounding box of the lesion in the markings and, subsequently, cropping the corresponding region from the original image, generating a new image that will be used in the subsequent steps of the method. Figure 3 illustrates the procedure for extracting the ROI.
With the ROIs extracted, the next step is image preprocessing.
3.3 ROI Preprocessing
Many studies have shown that image preprocessing has a significant impact on CNN results [9, 10, 20, 26]. In this step, we applied two filters sequentially to enhance the image representation.
Firstly, the anisotropic filter is applied, known for preserving edges in the image while effectively removing noise [30]. This filter is particularly advantageous in ultrasound (US) images, where edges are crucial for interpreting benign and malignant lesions. It preserves the structural characteristics of the image, such as edges and details, while adaptively reducing noise.
Next, the mean filter is applied. This filter is effective in removing background noise, smoothing the image, and enhancing the consistency of the structures identified by the anisotropic filter [18].
These filters combination enhances the visual US quality images and prepare them efficiently for feature extraction by CNNs, improving the accuracy of computer-aided diagnostic systems. Figure 4 illustrates the application of these filters.
We observed that the noise has been smoothed while the edges of the lesion have been preserved and enhanced. This process has proven to be effective in improving the training results of the Vision Transformer.
3.4 Balanced Data Augmentation
Before training the model, it is necessary to prepare the data by resizing and augmenting the minority class in the training dataset.
Firstly, it is important to note that after the ROI extraction step, the dimensions of the images in the dataset may vary in height and width. However, to optimize the training of specific CNN architectures, all images must have the same size in height and width. For this reason, we chose to resize all images to dimensions of (\(224\times {224}\)), which are suitable for training within the deep learning pipeline [19].
Next, observing 210 malignant images and 454 benign images, an imbalance problem between the classes becomes evident. Therefore, the database is divided into three sets: training, validation, and test. During training, data augmentation is performed based on the minority class (malignant images) to balance the number of samples between the classes.
Essentially, we compared the size of the minority class to that of the majority class. A set of hyperparameters is defined for data augmentation (up to a 5-degree rotation range, 10% image zoom-in, horizontal and vertical flipping). Thus, while the size of the minority class is smaller than that of the majority class, these transformations can occur randomly until the size of the majority class is reached. As a result, we increase the data and its representation.
After balancing the training samples, the Vision Transformer (ViT) architecture is defined and trained.
3.5 Vision Transformer Training
After preparing the data, the next step is to classify benign and malignant lesions in breast US images using the ViT-Base model [13], as shown in Fig. 5. First, the training and validation data were used to build the model. After constructing the model, the test dataset was employed to evaluate its robustness and performance.
The ViT-Base architecture includes an Embedding layer, multiple Transformer blocks, and a Multilayer Perceptron (MLP). The Embedding layer converts a 2D image into flattened token sequences while retaining positional information to feed into the stacked Transformer blocks. The standard Transformer encoder features multi-head self-attention layers interspersed with MLP blocks. Additionally, each block begins with a LayerNorm and includes residual connections at the end. The final MLP layer classifies the samples based on the output from the stacked Transformer blocks [13].
To facilitate training, pre-trained models (with the ImageNet database) were used with the default settings of the Keras library [4].
3.6 Evaluation Metrics
After training, we evaluate the model predictions using validation metrics to determine the effectiveness and robustness of the proposed method. In this study, the following metrics were employed: Accuracy (Acc), Sensitivity (Sens), Specificity (Spec), Precision (Prec), F1-score, and Area under the ROC curve (AUC-ROC) [14].
4 Results and Discussion
Now, after the method, the next step is to evaluate each technique that comprises it. In this section, we will present the results achieved by the proposed method.
To develop the method and experiments, a computer with the following specifications was used: Intel® Core™ i7 CPU@ 2.90GHz, Nvidia RTX-3060 12GB GPU, 16GB of RAM, and Windows 11 Pro operating system. The implementation was carried out in Python, using libraries such as Pandas, OpenCV, Scikit-Learn, and Keras.
The BUSI Dataset was divided into training (60%), validation (20%), and test (20%) sets. The training and validation sets were used to create the best model, while the test set was used for model evaluation. All hyperparameters used in ViT were defaulted and available in the Keras library.
To evaluate each step of the method, we conducted comparisons between the inclusion and exclusion of each proposed technique. The following sections present and discuss each of the experiments. Finally, we compare the results obtained with the literature presented in Sect. 2.
4.1 ROI Cropping Result
Two tests were conducted to evaluate the need for ROI cropping in the exams. First, the ViT (as described in Sect. 3.5) was executed on the original images (without the ROI). Then, the images were subjected to ROI cropping and processed again by ViT. Table 1 detailed ROI cropping results.
We can observe that by performing ROI cropping, there is a significant improvement in the Accuracy, Specificity, Precision, F1-score, and AUC metrics. Although the Sensitivity metric shows a reduction, it is important to highlight that the Accuracy and F1-score metrics, which provide an overall average of the models and a harmonic mean between Precision and Sensitivity, respectively, showed noteworthy improvements in results. It suggests that the model was able to enhance its classification capability.
4.2 ROI Preprocessing Result
After defining the ROI cropping step, the next step involves applying image preprocessing techniques to enhance its internal characteristics. To evaluate this step, we compared the results obtained without and with preprocessing (Table 2).
The results indicate that using the preprocessing step with anisotropic and mean filters resulted in improvements across all metrics. We highlight sensitivity, which had previously decreased with the ROI cropping step but has now exceeded the previous metric. Additionally, the F1-score deserves attention, showing a 6% improvement with the image enhancement step.
These improvements can be attributed to the ability of these techniques to enhance relevant features in the breast US images. For instance, the anisotropic filter effectively preserved edges while reducing noise, which is crucial for detecting image details. Similarly, the mean filter reduced noise impact, making it easier to identify subtle patterns.
Therefore, the effectiveness of these techniques in improving evaluation metrics can be attributed to their ability to optimize the representation of relevant features in the images, allowing the model to more accurately focus on important aspects during the classification process.
4.3 Balanced Data Augmentation Result
At this step, the training images of the malignant class undergo random data augmentation operations until they reach the same quantity as the majority class (benign class). This approach is crucial to increase the representativeness of the minority class, making the model more robust and balanced. Table 3 describes the results of the proposed technique in this step, comparing it with the previous step (without using data augmentation).
By employing the proposed data augmentation technique, we observed a significant improvement in results. All metrics showed notable enhancements, particularly sensitivity, which reached 99.25%. This increase in sensitivity indicates the model’s ability to correctly identify images from the malignant class. Furthermore, when observing the other metrics, all are above 97%.
We believe that the overall performance improvement of the model is a consequence of the increased diversity in the training data provided by data augmentation. By exposing the model to a greater variety of examples, a more comprehensive understanding of relevant characteristics is obtained, making it more capable of generalizing to new examples and handling variability in real-world data. It demonstrates that achieving greater diversity in the training set contributes to robust model performance.
4.4 ViT Result
To evaluate the performance of ViT, we compared it with two other widely used CNNs: ResNet50 [22] and EfficientNetB0 [29]. Overall, we kept all the steps of the method the same, replacing only ViT with the mentioned networks. It is worth noting that these networks are also pre-trained on ImageNet with default settings from the Keras library. Table 4 describes the results achieved.
It’s worth noting that all networks achieved noteworthy metrics for breast cancer classification in breast US images. It demonstrates that the steps before the CNN were crucial in improving the model’s performance. However, when analyzing the results achieved by ViT, a significant improvement of 2% to 6% is observed compared to the CNNs, surpassing them.
Transformers such as ViT distinguish themselves from traditional CNNs like ResNet50 and EfficientNetB0 by employing global attention mechanisms instead of convolution operations. It allows ViT to process relationships between all parts of the image simultaneously, capturing correlations on a broader scale. The architecture also excels in capturing long-range dependencies in the data, which is crucial for tasks where subtle patterns may occur in different parts of the image and require a global approach for accurate classification [13]. These factors contributed to its superior performance in breast cancer classification tasks in US images.
Therefore, we emphasize that the proposed method is robust in breast cancer classification. This method, used in CAD/CADx systems, can contribute to the rapid and effective diagnosis of this pathology, assisting specialist doctors in their daily clinical routine and thereby improving the quality of life for patients.
4.5 Related Works Comparison
To validate the proposed method concerning the studies found in the literature, we present Table 5, listing the studies from Sect. 2 along with the metrics achieved by them and by our work. It is important to note that, due to the methodological variations among the studies, the comparison is a reference for general trends and differences rather than providing an absolute measure of superiority.
The proposed method excels in all evaluated metrics, surpassing the results of the related works presented. Particularly, the F1-score and sensitivity metrics demonstrate the robustness of the proposed method, reflecting its ability to balance precision and sensitivity in detecting positive cases of breast cancer. These results reinforce the efficiency and reliability of the method, establishing it as a solid and effective approach for breast cancer classification in US images.
It is important to note that direct comparisons can be complex, as methods are developed on different datasets and with distinct approaches. However, upon examining the techniques employed, our work stands out as the only one to propose enhancements to ultrasound images using advanced image processing techniques. Additionally, we introduce automated data augmentation to improve model generalization. Furthermore, we innovate by using a Transformer-based CNN, an approach not addressed in related studies.
Thus, these approaches’ integration harmonizes classification techniques and surpasses the metrics of all other works. Therefore, we believe our work represents a significant contribution to the literature, holding a prominent position.
4.6 Case Study
In this section, we will analyze two case studies. In the first study, a lesion initially considered benign was incorrectly classified as malignant (Fig. 6 (A)). In the second case, a malignant lesion was erroneously classified as benign (Fig. 6 (B)).
In Fig. 6 (A), it can be observed that although the lesion is benign, its edges exhibit irregularities. This particularity may have contributed to the model’s misclassification as malignant. On the other hand, upon examining Fig. 6 (B), it is noticeable that the edges are more uniform, a common characteristic in benign lesions, which may have resulted in the incorrect classification of the lesion. However, it is to emphasize that these models can complement medical expertise, offering an additional tool for analysis and diagnosis. By combining them with careful clinical assessment and other relevant data, it is possible to improve the accuracy of diagnosis and thus play a crucial role in the fight against breast cancer.
5 Conclusion
For this work, we propose anisotropic and mean filters, balanced data augmentation, and the ViT network for breast cancer classification in ultrasound images. The filters proved effective in enhancing relevant image features, while balanced data augmentation contributed to improving the model’s robustness by balancing the representation of benign and malignant classes. Furthermore, the ViT network demonstrated high efficacy in classifying medical ultrasound images. The combination of these techniques resulted in promising outcomes, surpassing existing methods in the literature. Additionally, all validation metrics were above 97%, confirming the effectiveness of the proposed techniques.
In future work, we suggest investigating other preprocessing filters to validate the choice of anisotropic and mean filters. Additionally, another improvement to the method would be the optimization of the ViT network’s hyperparameters, as we used only the standard hyperparameters from the literature for this work.
Thus, we highlight that the proposed method is promising, potentially facilitating the clinical routine of specialist physicians and improving the chances of cure and survival for patients.
References
Al-Dhabyani, W., Gomaa, M., Khaled, H., Fahmy, A.: Dataset of breast ultrasound images. Data Brief 28, 104863 (2020)
Cao, Z., Yang, G., Chen, Q., Chen, X., Lv, F.: Breast tumor classification through learning from noisy labeled ultrasound images. Med. Phys. 47(3), 1048–1057 (2020)
Cheng, H.D., Shan, J., Ju, W., Guo, Y., Zhang, L.: Automated breast cancer detection and classification using ultrasound images: a survey. Pattern Recogn. 43(1), 299–317 (2010)
Chollet, F., et al.: Keras: the Python deep learning library. Astrophysics Source Code Library (2018)
Chougrad, H., Zouaki, H., Alheyane, O.: Deep convolutional neural networks for breast cancer screening. Comput. Methods Programs Biomed. 157, 19–30 (2018)
Diniz, J.O., Ferreira, J.L., da Silva, G.L., Quintanilha, D.B., Silva, A.C., Paiva, A.: Segmentação de coração em tomografias computadorizadas utilizando atlas probabilístico e redes neurais convolucionais. In: Anais do XXI Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 83–94. SBC (2021)
Diniz, J.O.B., et al.: Heart segmentation in planning CT using 2.5D U-Net++ with attention gate. Comput. Methods Biomech. Biomed. Eng. Imag. Vis. 11(3), 317–325 (2023)
Diniz, J.O.B., Diniz, P.H.B., Valente, T.L.A., Silva, A.C., de Paiva, A.C., Gattass, M.: Detection of mass regions in mammograms by bilateral analysis adapted to breast density using similarity indexes and convolutional neural networks. Comput. Methods Programs Biomed. 156, 191–207 (2018)
Diniz, J.O.B., Ferreira, J.L., Cortes, O.A.C., Silva, A.C., de Paiva, A.C.: An automatic approach for heart segmentation in CT scans through image processing techniques and Concat-U-Net. Expert Syst. Appl. 196, 116632 (2022)
Diniz, J.O.B., Ferreira, J.L., Diniz, P.H.B., Silva, A.C., de Paiva, A.C.: Esophagus segmentation from planning CT images using an atlas-based deep learning approach. Comput. Methods Programs Biomed. 197, 105685 (2020)
Diniz, J.O.B., Ferreira, J.L., Diniz, P.H.B., Silva, A.C., Paiva, A.C.: A deep learning method with residual blocks for automatic spinal cord segmentation in planning CT. Biomed. Sig. Process. Control 71, 103074 (2022)
Diniz, J.O., Quintanilha, D.B., de Carvalho Filho, A.O., Gomes Jr, D.L., Silva, A.C., Braz Jr, G., de Paiva, A.C., Luz, D.D.S.: Detecção de Covid-19 em imagens de raio-x de tórax através de seleção automática de pré-processamento e de rede neural convolucional. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 162–173. SBC, Porto Alegre, RS, Brasil (2023). https://doi.org/10.5753/sbcas.2023.229576
Dosovitskiy, A., et al.: An image is worth 16x16 words: Transformers for image recognition at scale. arXiv preprint arXiv:2010.11929 (2020)
Duda, R.: Pattern Classification and Scene Analysis. Wiley-Interscience Publication, vol. 512 (1973)
Eroğlu, Y., Yildirim, M., Çinar, A.: Convolutional neural networks based classification of breast ultrasonography images by hybrid method with respect to benign, malignant, and normal using mRMR. Comput. Biol. Med. 133, 104407 (2021)
Fernandes, A.G.S., Braz Junior, G., Diniz, J.O.B., Silva, A.C., Matos, C.E.F.: EfficientDeepLab for automated trachea segmentation on medical images. In: Brazilian Conference on Intelligent Systems, pp. 154–166. Springer (2023)
Figueredo, W., et al.: Abordagem computacional baseada em deep learning para o diagnóstico de endometriose profunda através de imagens de ressonância magnética. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 138–149. SBC, Porto Alegre, RS, Brasil (2023). https://doi.org/10.5753/sbcas.2023.229567
Gonzalez, R., Woods, R.: Digital Image Processing. Prentice Hall, Pearson (2008)
Júnior, D.A.D., et al.: Automatic method for classifying COVID-19 patients based on chest X-ray images, using deep features and PSO-optimized XGBoost. Expert Syst. Appl. 183, 115452 (2021)
Júnior, D.A.D., da Cruz, L.B., Diniz, J.O., Júnior, G.B., Silva, A.C.: Classificação automática de glóbulos brancos usando descritores de forma e textura e extreme gradient boosting. In: Anais do XXI Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 95–106. SBC, Porto Alegre, RS, Brasil (2021). https://doi.org/10.5753/sbcas.2021.16056
Kılıçarslan, G., Koç, C., Özyurt, F., Gül, Y.: Breast lesion classification using features fusion and selection of ensemble ResNet method. Int. J. Imaging Syst. Technol. 33(5), 1779–1795 (2023)
Koonce, B., Koonce, B.: ResNet 50. In: Convolutional Neural Networks with Swift for Tensorflow: Image Recognition and Dataset Categorization, pp. 63–72 (2021)
Marin, L.F., Silva, A.C., Bandeira, J.O.: Meta-data construction for selection of breast tissue biopsy slides image classifier to identify ductal carcinoma. In: 2019 8th Brazilian Conference on Intelligent Systems (BRACIS), pp. 729–734. IEEE (2019)
Moon, W.K., Lee, Y.W., Ke, H.H., Lee, S.H., Huang, C.S., Chang, R.F.: Computer-aided diagnosis of breast ultrasound images using ensemble learning from convolutional neural networks. Comput. Methods Programs Biomed. 190, 105361 (2020)
Pang, T., Wong, J.H.D., Ng, W.L., Chan, C.S.: Semi-supervised GAN-based radiomics model for data augmentation in breast ultrasound mass classification. Comput. Methods Programs Biomed. 203, 106018 (2021)
Santos, P., Brito, V., Filho, A.C., Sousa, A., Diniz, J., Luz, D.: Efficientbacillus: uma arquitetura profunda para detecção dos bacilos de koch. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 198–209. SBC, Porto Alegre, RS, Brasil (2023). https://doi.org/10.5753/sbcas.2023.229608
Shan, J., Alam, S.K., Garra, B., Zhang, Y., Ahmed, T.: Computer-aided diagnosis for breast ultrasound using computerized BI-RADS features and machine learning methods. Ultrasound Med. Biol. 42(4), 980–988 (2016)
Sirjani, N., et al.: A novel deep learning model for breast lesion classification using ultrasound images: a multicenter data evaluation. Phys. Med. 107, 102560 (2023)
Tan, M., Le, Q.: EfficientNet: rethinking model scaling for convolutional neural networks. In: International Conference on Machine Learning, p. 6105. PMLR (2019)
Weickert, J., et al.: Anisotropic Diffusion in Image Processing, vol. 1. Teubner Stuttgart (1998)
Zhuang, Z., Yang, Z., Raj, A.N.J., Wei, C., Jin, P., Zhuang, S.: Breast ultrasound tumor image classification using image decomposition and fusion based on adaptive multi-model spatial feature fusion. Comput. Methods Programs Biomed. 208, 106221 (2021)
Acknowledgments
This work was supported by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior - Brasil (CAPES) - Finance Code 001, Fundação de Amparo à Pesquisa do Maranhão (FAPEMA), Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), and Empresa Brasileira de Serviços Hospitalares (Ebserh) Brazil (Proc. 409593/2021-4). We also acknowledge LLM use for spell-checking, grammar correction, and assistance in translating specific terms.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2025 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Diniz, J.O.B. et al. (2025). AnisotropicBreast-ViT: Breast Cancer Classification in Ultrasound Images Using Anisotropic Filtering and Vision Transformer. In: Paes, A., Verri, F.A.N. (eds) Intelligent Systems. BRACIS 2024. Lecture Notes in Computer Science(), vol 15414. Springer, Cham. https://doi.org/10.1007/978-3-031-79035-5_7
Download citation
DOI: https://doi.org/10.1007/978-3-031-79035-5_7
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-79034-8
Online ISBN: 978-3-031-79035-5
eBook Packages: Computer ScienceComputer Science (R0)