Abstract
Colorectal cancer (CRC) is the second most prevalent type of cancer among men and women in Brazil, accounting for 9.1% of cancers in men and 9.2% in women between 2020 and 2022. CRC diagnosis typically relies on histopathological images, which can be challenging to interpret and prone to errors. This paper proposes a method for diagnosing CRC using histopathological images. This method involves enhancing images with MIRNet and classifying them as benign or malignant using InceptionV3. The results surpass the state-of-the-art by introducing, for the first time, enhancement through MIRNet, achieving an accuracy of 99.67% and an AUC of 0.9999. We believe that this method, combined with medical expertise, holds great promise for early CRC diagnosis.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Cancer encompasses diseases in which abnormal cells develop in the body due to random mutations. Among these conditions, colorectal cancer (CRC) is a significant contributor to morbidity and mortality [3]. In Brazil, approximately 45,630 new cases of CRC are estimated annually, with a higher incidence in men compared to women [21]. Globally, CRC accounted for over 1.9 million new cases in 2020, ranking it as the third most incident tumor among all cancers. The highest incidence rates of CRC were observed in men in Central, Northern, and Southern Europe, while among women, the highest incidences were found in Oceania and Northern Europe [3].
Colorectal cancer stands out as one of the most prevalent cancers in Brazil and worldwide, emphasizing the necessity to address detection and diagnostic methodologies that facilitate early identification [7]. Currently, specialists rely on visual analyses of histopathological image samples, a subjective process dependent on personal interpretation and pathologist experience [1]. The development of computational tools to support specialists and provide a second opinion is crucial for aiding in early diagnosis.
Artificial intelligence (AI) has made significant advancements, offering various viable diagnostic approaches [12, 13, 15, 17, 19, 23]. Once trained, it can assist specialists in quickly and effectively diagnosing CRC. Early diagnosis plays a fundamental role in treatment and patient survival. With recent advances in image processing and machine learning, AI techniques have shown promise in assisting in CRC diagnosis.
Therefore, this study presents a method for diagnosing CRC histopathological images using convolutional neural networks (CNNs) for image enhancement and classification between benign and malignant. Thus, we believe that the proposed method brings the following contributions:
-
Introduces the use of the MIRNet CNN, for the first time in the literature, to enhance the quality of histopathological images, improving diagnostic accuracy;
-
Utilizes another CNN, the InceptionV3, to classify the images pre-processed by MIRNet, achieving metrics superior to the literature.
Thus, the contributions mentioned suggest that the proposed method is an effective tool to assist specialists in an early diagnosis of CRC cancer, potentially playing a critical role in effective treatment and patient survival.
2 Related Works
Colorectal cancer has sparked extensive research into the development of computational methods to aid specialists in its classification. In this section, related works on the topic are presented.
Dabass et al. [8] highlight the importance of computational methods for CRC classification. They propose a supervised deep learning technique for five-grade cancer classification, achieving an accuracy of 96.97% for two-grade and 93.24% for five-grade classification.
Candelero et al. [5] propose an automated pathology detection method, focusing on CRC, breast cancer, and non-Hodgkin lymphomas. Their method combines feature extraction techniques such as multi-scale and multidimensional fractals, Haralick descriptors, and CNN for pattern recognition. Following feature selection, evolutionary and machine learning algorithms, including SVM, K*, and Random Forest, were applied for classification. The best combination resulted in an accuracy of 91.06% for CRC.
Mangal et al. [20] highlight the importance of accurately categorizing histopathological images of the colon. Using the same dataset of our work for validation, they achieved an impressive accuracy of 96.61% with a CNN architecture they proposed. On the other hand, Bukhari et al. [4] emphasize the precise histopathological diagnosis of colorectal cancer. Their approach involves utilizing images and CNNs. By training on ten thousand images and comparing three different CNNs (ResNet-18, ResNet-30, and ResNet-50), they identified ResNet-50 as the most accurate, achieving an outstanding 93.91% accuracy.
Hidayah et al. [18] introduce a conventional approach to classify colorectal cancer, employing texture analysis with GLRLM and k-fold cross-validation. This research attains an accuracy rate of 85.57%, a sensitivity of 91.72%, and a specificity of 80.55%. All two studies mentioned utilize the identical dataset as the proposed method. On the other hand, Swarna et al. [24] present two models for colon cell classification based on image data - one utilizing the InceptionV3 model, and the other combining predictions from three basic sequential CNN models. The authors achieve accuracy rates of 99.4% and 99.95%, respectively.
In a recent study conducted by Rajinikanth et al. [22], they introduced a new deep learning framework designed to categorize colon slides into either normal or cancerous classes. This methodology encompasses several steps, including resizing, preprocessing, extraction of deep features, and binary classification using 5-fold cross-validation. The research dataset comprises 4,000 images, evenly split between normal and cancerous samples. Their findings reveal that employing deep features leads to a remarkable 99% accuracy rate.
Table 1 describes the related works presented, along with the techniques and accuracy values.
We notice that in most studies, CNN usage stands out for CRC classification, displaying significant metrics, often surpassing 90%. Moreover, feature extraction, whether traditional or deep features, is also commonly employed for this purpose. However, there is a lack of emphasis on image enhancement. Enhancing images is crucial to highlight important features and optimize their representation for subsequent steps, including feature extraction or CNN utilization.
Addressing this gap, the proposed method introduces the use of a CNN called MIRNet for the automatic enhancement of CRC histopathological images. Furthermore, considering the results achieved in the literature, we propose utilizing InceptionV3 for classification after applying MIRNet [26], as it was the CNN that exhibited the best performance [24].
3 Materials and Method
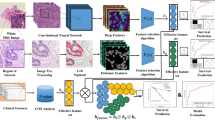
In this section, we describe the steps that compound the proposed method. The proposed method is divided into four steps: database acquisition; database pre-processing using MIRNet; InceptionV3 as a classifier between benign and malignant classes; and finally, a validation step to evaluate the proposed method. Figure 1 presents the mentioned steps.
3.1 Database
For this study, we turned to the extensive database available on Kaggle, titled Lung and Colon Cancer Histopathological Images. This dataset, a rich and diverse source, comprises five distinct classes, each containing 5,000 images. The categories include benign lung tissue, lung adenocarcinoma, lung squamous cell carcinoma, colon adenocarcinoma, and benign colon tissue [2].
For our method development, we concentrated on a database comprising 10,000 images associated with colon adenocarcinoma and benign colon tissue. Our analysis aims to discern the distinguishing features between benign and malignant images utilizing deep learning-based enhancement techniques. Figure 2 offers a visual representation of a sample from each class.
Additionally, the images have dimensions of \(768\times 768\) pixels. Thus, for the subsequent steps of the method, resizing to \(256\times 256\) was performed. According to [10, 16], this size is suitable for processing within the deep learning pipeline.
3.2 MIRNet Pre-Processing
Many works show that image preprocessing can make the results better [9, 11, 14]. In this step, the images are processed by MIRNet to assess the enhancement of the database. MIRNet (Multiple Iterative Reconstruction Network) is a deep neural network designed to perform image enhancement. It operates in multiple iterations, progressively refining the quality of the input images [26].
We highlight that it is the first time the MIRNet has been used in histopathological images of CRC as image pre-processing. Thus, image pre-processing techniques play a crucial role in improving results. It is not an easy task to select a set of the best pre-processing to be applied. This way, employing an AI-based technique that automatically enhances the image features increases the method’s performance. Figure 3 illustrates the architecture of MIRNet.
According to [26], MIRNet follows an iterative approach, where each iteration enhances the input image by learning hierarchical representations of image features. It reconstructs the image using learned residual information, which are differences between the input and reconstructed images. These residuals are used to refine the input image in each iteration, resulting in changes in texture patterns. Various configurations were tested to validate the method, as discussed in Sect. 4.
As shown in Fig. 2, histopathological images of CRC often exhibit variations in quality and appearance, which can hinder the accurate classification of tissue. Therefore, by opting to use MIRNet as preprocessing, it can highlight important features of CRC tissue that may not be easily discernible, such as morphological, structural, and textural characteristics relevant for distinguishing benign from malignant tissue.
3.3 InceptionV3 Training
With the images pre-processed by MIRNet, the next step is to classify them as malignant or benign. For this purpose, a CNN will be trained. Firstly, the data is split into training, validation, and test sets. It is worth noting that one of the characteristics of CNNs is the need for a large number of training samples to effectively learn the features during training.
At this step, data augmentation techniques were applied to increase the size of the training dataset. The techniques used included random rotation from -30 to 30\(^\circ \)C, width_shift_range and height_shift_range also from -30 to 30\(^\circ \)C and horizontal and vertical flipping operations. These operations were performed using the Keras library [6].
For training the CNN, the InceptionV3 was chosen. Upon reviewing the literature analysis (Sect. 2), this was the architecture that yielded the best results in CRC analysis. Moreover, InceptionV3 is capable of extracting complex and discriminative features from histopathological images of CRC. Its efficient architecture, which utilizes parallel convolutional modules called Inception modules, allows for a richer and more efficient representation of image features [25].
Furthermore, the transfer learning technique can be easily applied to InceptionV3 by leveraging its pre-trained weights and fine-tuning them to fit the specific data of the CRC classification problem. Therefore, the weights from ImageNet were utilized. By adjusting these weights to fit the specific data of the CRC classification problem, the network can learn more specialized and discriminative representations to distinguish between benign and malignant tissue.
Thus, training is performed by InceptionV3 with data augmentation and transfer learning from ImageNet, and validated on the validation set to find the best model. Finally, the test data is presented to the model, and validation metrics are extracted.
3.4 Validation Metrics
In computational analysis of histopathological images, accurate classification of samples between benign and malignant plays a fundamental role in aiding clinical diagnosis. Therefore, to evaluate the proposed method’s performance metrics frequently applied in the area of medical imaging and literature were used: accuracy, precision, sensitivity, specificity, F1-score, and area under the ROC curve (AUC). Calculate the metrics, it was based on the confusion matrix, which takes into account four variables: TP denotes true positives, TN represents true negatives, FP signifies false positives, and FN indicates false negatives.
where TPR (True Positive Rate) represents the rate of true positives; fpr (false Positive Rate) represents the rate of false positives. The integral is calculated from 0 to 1, representing the area under the ROC curve.
4 Results and Discussion
In this section, we present the results obtained through the application of the proposed method. Firstly, we describe the training environment used. Then, we detail the results achieved by the proposed method. Subsequently, a comparison with the literature and case studies is conducted. Finally, the advantages and limitations of the method are discussed.
4.1 Training Environment
To validate the proposed work, the following hardware resources were used: an Intel® Core™ i7 CPU 2.30GHz processor, an Nvidia® RTX-3070 8GB GPU, 16GB of RAM, and the Windows 11 Pro operating system.
TensorFlow and the Keras library were employed to build and analyze the proposed method. TensorFlow is a widely used library for developing CNNs and serves as the backend for Keras. Additionally, OpenCV was used for image manipulation, and Scikit-Learn to assist in various stages of the method, as well as to apply validation metrics.
4.2 Experiments
In the experiments, we sought to validate each step of the proposed method. Initially, the dataset was divided into training (80%) and testing (20%) sets, with a validation subset comprising 20% of the training set, as described in Table 2.
Next, pre-processing was applied using MIRNet. It is important to highlight that all hyperparameters used in MIRNet were the same as those described by [26]. For this purpose, four datasets were trained with 50, 100, 200, and 500 epochs, respectively. Figure 4 illustrates the result of some images for each of these sets.
We observed that for each dataset, different aspects of the images are enhanced, varying according to the number of epochs. Visually, the dataset trained with 50 epochs presents a visually harmonized result, while those trained with 100, 200, and 500 epochs lost some characteristics. This suggests the need for a more in-depth analysis of the metrics to verify if there was indeed an improvement in the pre-processing quality.
Subsequently, each dataset is submitted to InceptionV3. The hyperparameters used were those presented by [25], available in the Keras library. In addition, data augmentation and transfer learning steps were included, as described in Sect. 3.3. The models were trained for 50 epochs, with a batch size of 16, using the Adam optimizer and binary cross-entropy as loss.
Table 3 presents the results for training InceptionV3 for each dataset and the original images.
The analysis of the metrics reveals a significant improvement in the model’s performance after pre-processing with MIRNet compared to the original images. In percentage terms, we observe a considerable increase in all metrics across all datasets processed by MIRNet. When comparing the MIRNet datasets, we note that those trained with 50 and 100 epochs exhibit the best metrics. Specifically, the MIRNet dataset with 50 epochs stands out as the best, with 99.67% accuracy, 99.73% precision, 99.73% sensitivity, 99.70% specificity, 99.67% F1-score, and 0.9999 AUC. These results suggest that a shorter training of 50 epochs may be sufficient to achieve excellent performance.
These results underscore its significance in CRC diagnosis. The substantial improvement in performance metrics after pre-processing with MIRNet indicates that this approach can assist in more accurate identification of benign and malignant tissues in CRC histopathological images. The high metrics achieved with MIRNet suggest that this technique could be a valuable tool to aid pathologists and physicians in early CRC diagnosis and treatment, thereby contributing to better clinical outcomes and improved quality of life for patients.
4.3 Comparison with the Literature
Observing Sect. 2, our work stands out. Table 4 presents our work along with related works.
Though direct comparisons are challenging due to differences in datasets and methodologies, our proposed method has demonstrated high effectiveness in classifying histopathological images of CRC. Leveraging MIRNet and InceptionV3 techniques, our method achieved an accuracy of 99.67
Considering metrics beyond accuracy, our method also yielded promising results in terms of precision, sensitivity, specificity, F1-score, and AUC. These metrics provide a comprehensive view of our model’s performance, demonstrating its ability to successfully distinguish between benign and malignant tissue in CRC histopathological images.
4.4 Case Study
In this section, we will describe two case studies of the proposed method. In Fig. 5 (A), we present an example of a cancer image classified as benign, and in Fig. 5 (B), we have a benign image classified as cancer.
Visually, it can be observed that both images have very similar texture characteristics. The misclassification error may be attributed to a possible incorrect labeling or to the fact that, due to their similarity, MIRNet may have enhanced features that confused InceptionV3 in classification. However, it is essential to highlight that, despite the classification errors, the method demonstrated considerable metrics, suggesting a notable overall performance.
4.5 Discussion
Now, let’s discuss the advantages and limitations of our method for classifying CRC histopathological images.
-
Our proposed method automates the process of distinguishing histopathological images, reducing manual intervention and, combined with medical expertise, enhances diagnostic efficiency.
-
Our approach stands out by employing MIRNet for the first time in the context of CRC histopathological images. MIRNet automatically highlights relevant features, significantly contributing to the differentiation between malignant and benign tissues.
-
We propose the use of data augmentation and transfer learning during training, bringing effective techniques for model enhancement.
-
By adopting the InceptionV3 architecture for classification, our method benefits from the ability of this CNN to extract complex and discriminative features from images, resulting in superior performance.
-
Our method includes metrics such as accuracy, precision, sensitivity, specificity, F1-score, and AUC, providing a more comprehensive and robust evaluation of the model’s performance.
-
Due to the flexible architectures and comprehensive metrics used, our method has the potential to be applied in other areas of medicine and in the classification of different types of histopathological images.
However, it is worth noting that despite the method being robust, achieving 99% in all validation metrics, it does have some limitations:
-
While MIRNet is a powerful technique for image pre=processing, determining the best hyperparameters to optimize its performance automatically is still necessary.
-
We limited our analysis to the InceptionV3 architecture for classification, but exploring other CNN architectures could further validate the proposed method.
-
Although we conducted experiments to empirically adjust the model’s hyperparameters, an automated approach for selecting the best hyperparameters could enhance the method’s performance.
These considerations about advantages and limitations provide a comprehensive view of our method and highlight areas for future work.
5 Conclusion
The proposed method was developed to assist in the diagnosis of CRC, one of the leading causes of morbidity and mortality worldwide. We propose a robust automatic method for classifying CRC histopathological images, aiming to improve clinical diagnosis. The method represents a contribution by being the first to utilize MIRNet for the preprocessing of CRC histopathological images. Furthermore, with the use of the InceptionV3 architecture, we obtained promising results, demonstrating the model’s ability to extract discriminative features and achieve metrics superior to the literature.
As future work, other CNN architectures beyond InceptionV3 could be explored to validate their use and compare them with existing ones. Additionally, it is suggested to investigate automated techniques for hyperparameter selection, aiming to make the method even more robust.
References
Abdelsamea, M.M., Zidan, U., Senousy, Z., Gaber, M.M., Rakha, E., Ilyas, M.: A survey on artificial intelligence in histopathology image analysis. Wiley Interdisc. Rev. Data Min. Knowl. Discov. 12(6), e1474 (2022)
Borkowski, A.A., Bui, M.M., Thomas, L.B., Wilson, C.P., DeLand, L.A., Mastorides, S.M.: Lung and colon cancer histopathological image dataset (lc25000). arXiv preprint arXiv:1912.12142 (2019)
Bray, F., Ferlay, J., Soerjomataram, I., Siegel, R., Torre, L., Jemal, A., et al.: Erratum: global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 70(4), 313 (2020)
Bukhari, S.U.K., Syed, A., Bokhari, S.K.A., Hussain, S.S., Armaghan, S.U., Shah, S.S.H.: The histological diagnosis of colonic adenocarcinoma by applying partial self supervised learning. MedRxiv, pp. 2020–08 (2020)
Candelero, D., Roberto, G.F., Do Nascimento, M.Z., Rozendo, G.B., Neves, L.A.: Selection of CNN, Haralick and fractal features based on evolutionary algorithms for classification of histological images. In: 2020 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), pp. 2709–2716. IEEE (2020)
Chollet, F., et al.: Keras. https://keras.io (2015)
Crespo, J., Victorino, A.P., Araujo, K., Araujo, L.H., Vieira, F.M.D.A.C.: Colorectal cancer biomarkers and their impact on the clinical practice. Braz. J. Oncol. 17, 1–14 (2021)
Dabass, M., Vig, R., Vashisth, S.: Five-grade cancer classification of colon histology images via deep learning. In: Communication and Computing Systems, pp. 18–24. CRC Press (2019)
Diniz, J.O., et al.: Detecçao de COVID-19 em imagens de raio-x de tórax através de seleçao automática de pré-processamento e de rede neural convolucional. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 162–173. SBC (2023)
Diniz, J.O., et al.: Efficientxyz-deepfeatures: seleção de esquema de cor e arquitetura deep features na classificação de câncer de cólon em imagens histopatológicas. In: Simpósio Brasileiro de Computação Aplicada à Saúde (SBCAS), pp. 82–93. SBC (2024)
Diniz, J.O.B., et al.: Heart segmentation in planning CT using 2.5D U-Net++ with attention gate. Comput. Methods Biomech. Biomed. Eng. Imag. Vis. 11(3), 317–325 (2023)
Diniz, J.O.B., Diniz, P.H.B., Valente, T.L.A., Silva, A.C., Paiva, A.C.: Spinal cord detection in planning CT for radiotherapy through adaptive template matching, IMSLIC and convolutional neural networks. Comput. Methods Programs Biomed. 170, 53–67 (2019)
Diniz, J.O.B., Diniz, P.H.B., Valente, T.L.A., Silva, A.C., de Paiva, A.C., Gattass, M.: Detection of mass regions in mammograms by bilateral analysis adapted to breast density using similarity indexes and convolutional neural networks. Comput. Methods Programs Biomed. 156, 191–207 (2018)
Diniz, J.O.B., Ferreira, J.L., Cortes, O.A.C., Silva, A.C., de Paiva, A.C.: An automatic approach for heart segmentation in CT scans through image processing techniques and Concat-U-Net. Expert Syst. Appl. 196, 116632 (2022)
Diniz, J.O.B., Ferreira, J.L., Diniz, P.H.B., Silva, A.C., de Paiva, A.C.: Esophagus segmentation from planning CT images using an atlas-based deep learning approach. Comput. Methods Programs Biomed. 197, 105685 (2020)
Diniz, J., et al.: Detecção de COVID-19 em imagens de raio-x de tórax através de seleção automática de pré-processamento e de rede neural convolucional. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 162–173. SBC, Porto Alegre, RS, Brasil (2023). https://doi.org/10.5753/sbcas.2023.229576
Figueredo, W., et al.: Abordagem computacional baseada em deep learning para o diagnóstico de endometriose profunda através de imagens de ressonância magnética. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 138–149. SBC, Porto Alegre, RS, Brasil (2023). https://doi.org/10.5753/sbcas.2023.229567
Hidayah, N., Ramadanti, A.N., Novitasari, D.C.R.: Classification of colon cancer based on hispathological images using adaptive neuro fuzzy inference system (ANFIS). Khazanah Informatika Jurnal Ilmu Komputer dan Informatika 9(2), 162–168 (2023)
Júnior, D.D., Cruz, L., Diniz, J., Júnior, G.B., Silva, A.: Classificação automática de glóbulos brancos usando descritores de forma e textura e extreme gradient boosting. In: Anais do XXI Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 95–106. SBC, Porto Alegre, RS, Brasil (2021). https://doi.org/10.5753/sbcas.2021.16056
Mangal, S., Chaurasia, A., Khajanchi, A.: Convolution neural networks for diagnosing colon and lung cancer histopathological images. arXiv preprint arXiv:2009.03878 (2020)
de Oliveira Santos, M., de Lima, F.C.d.S., Martins, L.F.L., Oliveira, J.F.P., de Almeida, L.M., de Camargo Cancela, M.: Estimativa de incidência de câncer no brasil, 2023-2025. Revista Brasileira de Cancerologia 69(1) (2023)
Rajinikanth, V., Kadry, S., Mohan, R., Rama, A., Khan, M.A., Kim, J.: Colon histology slide classification with deep-learning framework using individual and fused features. Math. Biosci. Eng. 20(11), 19454–19467 (2023)
Santos, P., Brito, V., Filho, A.C., Sousa, A., Diniz, J., Luz, D.: Efficientbacillus: uma arquitetura profunda para detecção dos bacilos de koch. In: Anais do XXIII Simpósio Brasileiro de Computação Aplicada à Saúde, pp. 198–209. SBC, Porto Alegre, RS, Brasil (2023). https://doi.org/10.5753/sbcas.2023.229608
Swarna, I.J., Hashi, E.K.: Detection of colon cancer using inception v3 and ensembled CNN model. In: 2023 International Conference on Electrical, Computer and Communication Engineering (ECCE), pp. 1–6. IEEE (2023)
Szegedy, C., Vanhoucke, V., Ioffe, S., Shlens, J., Wojna, Z.: Rethinking the inception architecture for computer vision. In: Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, pp. 2818–2826 (2016)
Zamir, S.W., et al.: Learning enriched features for real image restoration and enhancement. In: Vedaldi, A., Bischof, H., Brox, T., Frahm, J.-M. (eds.) ECCV 2020. LNCS, vol. 12370, pp. 492–511. Springer, Cham (2020). https://doi.org/10.1007/978-3-030-58595-2_30
Acknowledgments
To the support of the Coordination for the Improvement of Higher Education Personnel (CAPES) - Financing Code 001; the Foundation for Research Support of Maranhão (FAPEMA); the National Council for Scientific and Technological Development (CNPq); and the Brazilian Company of Hospital Services (Ebserh) - Brazil (Proc. 409593/2021-4).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2025 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
Ribeiro, N.P. et al. (2025). Improving Colorectal Cancer Diagnosis Using MIRNet and InceptionV3 on Histopathological Images. In: Paes, A., Verri, F.A.N. (eds) Intelligent Systems. BRACIS 2024. Lecture Notes in Computer Science(), vol 15414. Springer, Cham. https://doi.org/10.1007/978-3-031-79035-5_22
Download citation
DOI: https://doi.org/10.1007/978-3-031-79035-5_22
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-79034-8
Online ISBN: 978-3-031-79035-5
eBook Packages: Computer ScienceComputer Science (R0)