Abstract
In this study, we developed and evaluated two medical large language models, Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2, specifically designed for Brazilian Portuguese. Utilizing the Low-Rank Adaptation (LoRA) technique, our models achieved significant improvements in generating synthetic clinical text, particularly in terms of Authenticity of Format and Structure, Spelling Accuracy, and Clinical Coherence. The evaluation, conducted by medical students using a 5-point Likert scale, demonstrated the effectiveness of our approach compared to baseline models. The scores indicate superior performance compared to baseline models such as LlaMA-2-7B and Mistral-7B-v0.2. Our results suggest that these resource-efficient models can effectively generate clinically relevant text, maintaining high standards of structure, accuracy, and coherence. Future work will focus on expanding datasets, refining evaluation protocols, and enhancing model robustness to further improve performance across various medical tasks.
Access provided by University of Notre Dame Hesburgh Library. Download conference paper PDF
Similar content being viewed by others
1 Introduction
Foundation models have gained significant attention for their strong ability to process various data types, particularly text. Large Language Models (LLMs), a subset of these architectures, are central in natural language processing (NLP) research due to their capability to generate coherent and contextually relevant text. This makes them instrumental in applications from automated content creation to conversational agents. The versatility and scalability of LLMs highlight their potential to revolutionize interactions with digital information, driving advancements in AI.
In the medical field, LLMs enhance the efficiency and accuracy of clinical decision-making processes [5]. These models excel in interpreting complex medical language and extracting relevant information from unstructured texts [29], aiding in the understanding of patient histories and treatment outcomes [29]. LLMs streamline administrative procedures and support diagnostic/therapeutic decisions [28], potentially improving patient outcomes and operational efficiencies [5]. For example, LLMs like Med-PaLM can achieve passing scores on medical licensing exams, showcasing their potential in clinical knowledge and question-answering tasks [29]. They also extract structured information from clinical notes and reports in various languages, enhancing diagnostic accuracy and aiding in health-related information dissemination [28].
The lack of resources in languages other than English is a persistent issue in NLP research, and the development of LLMs is no exception. This gap is further worsened by the substantial computational power required to train LLMs from scratch, a resource predominantly accessible to large corporations. It is essential to find ways to make these technologies accessible to the research and academic community by leveraging publicly available multilingual models and adapting them to specific contexts.
In this work, we focus on creating a useful Medical LLM for the Brazilian Portuguese language (pt-BR) using minimal computational resources. Our methodology involved continuing the pre-training (or unsupervised fine-tuning) of LlaMA-2-7b and Mistral-7b-v0.2 base models using three different datasets of clinical narratives. We applied the LoRA (Low-Rank Adaptation) technique to enhance the efficiency of this process. The new models, called Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2, were evaluated on their ability to generate synthetic clinical data, and results showed that our models outperformed other baseline models in generating clinically relevant data.
This initiative is part of a collaborative project between HAILab and Comsentimento, named MED-LLM-BRFootnote 1, which aims to develop multiple medical LLMs for pt-BR, including base models and task-specific models, with different sizes. To the best of our knowledge, this is the first clinical LLM model publicly available for the Portuguese language.
Our work addresses the specific needs and gaps in the Brazilian Health Informatics community, providing a valuable resource for clinicians and researchers to utilize in their healthcare AI projects. By overcoming linguistic and technical challenges, we contribute to the broader field of natural language processing and medical informatics, highlighting the significance of domain-specific knowledge in medical language processing.
2 Related Work
The development of Large Language Models (LLMs) has revolutionized natural language processing (NLP), especially in the medical field. This section reviews advancements in Transformer architectures, the importance of transfer learning and fine-tuning, and computational strategies for optimizing these models. We examine key initiatives in other languages, particularly Portuguese, and highlight prominent medical LLMs to provide context for our resource-efficient Medical LLM for Brazilian Portuguese.
The advent of Transformers, introduced by Vaswani et al. [26], revolutionized the field of natural language processing (NLP). This architecture uses self-attention mechanisms to handle long-range dependencies in text, significantly improving performance over previous recurrent neural network (RNN) and convolutional neural network (CNN) models.
Transformers can be configured as encoder or decoder models, supporting both generative and non-generative tasks. Encoder models, like BERT [7], focus on understanding and encoding input text into a dense representation. Decoder models, such as GPT-3 [3], generate text based on an input sequence. Encoder-decoder models, like the original Transformer [26], combine both functionalities and are effective for tasks like translation and summarization.
In the context of transfer learning, it’s crucial to distinguish between base models and task-specific models. Base models, like BERT and GPT-3, are pre-trained on large corpora to learn general language representations. These models can then be fine-tuned on specific tasks, such as sentiment analysis or named entity recognition, to create task-specific models that retain the general language understanding while being optimized for particular tasks.
Training and fine-tuning large language models pose significant computational challenges. Parameter Efficient Fine-Tuning (PEFT) methods, like Low-Rank Adaptation (LoRA) [11], introduce trainable low-rank matrices into each layer of a pre-trained model, reducing trainable parameters while maintaining performance and lowering costs. Models like LlaMA 2 [25] and Mistral [13] are designed for efficiency in training and inference. LlaMA 2 and Mistral 7B models offer superior performance and resource efficiency, with LlaMA 2 excelling in benchmarks and being accessible for various applications due to its smaller size. Mistral 7B’s Mixtral architecture uses expert mixture techniques to enhance performance and scalability, making high-performance LLMs more accessible and practical for specialized tasks.
By utilizing PEFT methods, it is possible to adapt large models like those aimed at pt-BR more efficiently, enabling broader application despite computational constraints. These techniques are crucial for developing specialized models in resource-constrained environments, ensuring that advancements in NLP are accessible and applicable across different languages and domains.
While most large language models have been developed for English, significant efforts have been made to create models for other languages. For example, mBERT (multilingual BERT) handles 104 languages by training on a multilingual corpus [27], and XLM-R [6] extends this idea by pre-training on a massive multilingual dataset.
There are initiatives to develop models for pt-BR, such as Sabiá [20], Sabiá-2 [1] and Bode [9]. However, relatively few models exist for this language, despite the large portuguese-speaking population (almost 250 million speakers).
In the medical field, models like BioBERT [16] and ClinicalBERT [12] are fine-tuned on domain-specific corpora to capture the unique terminology and context of medical texts. Med-BERT [21], trained on electronic health records (EHRs), predicts patient outcomes and assists in clinical decision-making.
Meditron and BioMistral have shown significant performance gains by extending pre-training on curated medical corpora [4, 15]. GatorTronGPT, trained from scratch using 277 billion words of mixed clinical and English text with a GPT-3 architecture, improves biomedical NLP for medical research [19]. OpenBioLLM-70B leverages the Meta-Llama-3-70B-Instruct model to achieve state-of-the-art performance on various biomedical tasks [2].
In the pt-BR medical context, the two main resources are the BioBERTpt [23] and CardioBERTpt [22]. BioBERTpt is a biomedical LLM, fine-tuned from multilingual BERT using clinical and biomedical data, showing promising results in clinical text tasks like Named Entity Recognition and Negation detection. While CardioBERTpt was fine-tuned on clinical data specifically from Cardiology. The only available generative model for pt-BR is GPT2-bio-pt [24], which was trained exclusively on biomedical data from medical articles. The model has several limitations such as to be based on a model trained on automatically translated text, absence of clinical narratives knowledge and small context window (512 tokens).
3 Method
This chapter describes our methodology divided into three key steps: Data Acquisition, Model Architecture and Training, and Experimental Setup. Each subsection details the processes and techniques employed to build and evaluate our models.
3.1 Data Acquisition
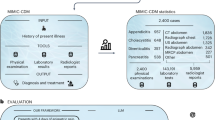
Our project combined data from three distinct clinical datasets, totaling 2.4GB of text and 309 151 121 tokens. The first one comes from the same data sources used in the SemClinBr project [18], which are composed of 2 100 546 clinical narrative entries from multiple Brazilian Hospitals. The dataset contains diverse document types (e.g., discharge summaries, ambulatory notes, nursing notes) and medical specialties (e.g., cardiology, nephrology, endocrinology). The electronic health record (EHR) data used in the study were de-identified and approved by the PUCPR Research Ethics Committee, certificate of presentation for ethical appreciation number 51376015.4.0000.0020.
The BRATECA dataset [8] was collected as well, and it is composed of 73 040 admission notes from 10 Brazilian Hospitals and associated with multiple medical departments (e.g., obstetrics, surgery, emergency, COVID-19, intensive care, ambulatory). All the data was anonymized and granted ethical approval by the National Research Ethics Committee under the number 46652521.9.0000.5530. The dataset is available under PhysioNet Credentialed Health Data Use [10].
Finally, we utilized the data used in Lopes et al. work [17], which consists mostly of neurology clinical cases collected from medical journals written in European Portuguese. The dataset contains 3678 medical texts and it is publicly availableFootnote 2.
3.2 Model Architecture and Training
In this study, we fine-tuned the LlaMA-2 and Mistral base models using the LlaMA-Factory framework [30], focusing on 7 billion parameter (7B) models to minimize computational resources. The models were trained over 2 epochs with a learning rate of 2e−5, using Low-Rank Adaptation (LoRA) for efficient weight updates. LoRA introduces trainable low-rank matrices into each layer, reducing trainable parameters and GPU memory requirements.
To further optimize memory and computational usage, we applied LoRA with 16-bit precision on the q_proj and v_proj projections, setting LoRA R to 8, LoRA Alpha to 16, and LoRA Dropout to 0.1. We used the AdamW optimizer with settings \(\beta _1 = 0.9\) and \(\beta _2 = 0.999\) to balance rapid convergence with stability during training. The main difference between the two fine-tuned models was the max_position_embeddings parameter configuration: 4096 for LlaMA-2 and 32768 for Mistral.
Training was conducted on Google Cloud Platform (GCP) with an NVIDIA Tesla A100 GPU, 12 vCPUs, and 85 GB of RAM. The resulting models, Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2, had their Training Loss shown in Fig. 1. We used 2.4 GB of clinical data for this study. Training the Clinical-BR-LlaMA-2-7B model involved 309,151,121 tokens and took 47 h and 56 min, costing R$ 1250.94. The Clinical-BR-Mistral-7B-v0.2 model took 50 h and 18 min, costing R$ 1202.73.
3.3 Experimental Setup
The models trained in this work are base models, trained unsupervised on a text corpus to learn clinical language representations. They serve as foundations for further fine-tuning for specific tasks. Evaluating base models typically involves benchmarks for tasks like language understanding, text generation, and question answering. However, these benchmarks can be imprecise, often relying on standard metrics that may not capture the nuances of language or generalization to unseen data. They might not reflect real-world performance, especially in specialized fields like healthcare, where domain-specific knowledge is crucial. For instance, most medical LLMs are evaluated on English Question Answering benchmarks like MedQA [14], which do not measure the model’s ability to interpret clinical data in electronic medical records, the focus of this work.
To overcome limitations, we used the models to generate synthetic clinical text and evaluated their performance on three criteria using a 5-point Likert scale (Fig. 2): Authenticity of Format and Structure, Spelling Accuracy, and Clinical Coherence.
Criterion 1 Authenticity of Format and Structure: Evaluated if the model’s output adhered to clinical document norms, including layout and sectioning. The highest score was given to models that perfectly matched the standard.
Criterion 2 Spelling Accuracy: Focused on linguistic correctness, including spelling and medical terminology in pt-BR. Models lost points for incorrect grammar or generating text in a different language.
Criterion 3 Clinical Coherence: Assessed if the content logically correlated with the clinical history, focusing on relevance and precision. This criterion involved more subjectivity due to varying evaluator backgrounds.
We selected 100 random clinical notes, not part of the training corpus, and used excerpts as input for five models to complete the notes. This aimed to measure the models’ proficiency in producing consistent clinical notes. Medical students with experience in clinical notes evaluated the generated texts from our models (Clinical-BR-LlaMA-2-7B, Clinical-BR-Mistral-7B-v0.2) and three reference models (LlaMA-2-7B, Mistral-7B-v0.2, Sabia-7B). They assigned scores from 1 to 5 on a Likert scale for each of the three criteria. The evaluation was blind; students did not know which model generated each note.
LlaMA-2-7B and Mistral-7B-v0.2 served as benchmarks to measure improvements from our training. Sabia-7B, a Portuguese-trained model, was included for comparison. This selection allowed a thorough performance analysis across benchmarks. GPT2-bio-pt was excluded due to its limited context size, which is inadequate for clinical text generation.
4 Results
This section presents the evaluation outcomes of Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2 models, alongside three baselines: LlaMA-2-7B, Mistral-7B-v0.2, and Sabia-7B. The evaluation focused on Authenticity of Format and Structure, Spelling Accuracy, and Clinical Coherence, using a 5-point Likert scale. We analyze the average scores for each criterion, the frequency each criterion was met, and perform an error analysis. This helps us understand the strengths and weaknesses of each model, highlight improvements in our fine-tuned models, and identify areas for future enhancement.
4.1 Clinical Text Generation
The average score of the models for each criterion is presented in Table 1 and stacked in Fig. 3. In addition, Table 2 presents the frequencies of score for each criterion in the evaluation of each model. In Table 3, we can see some examples of clinical notes generated with our models that were successful in the three evaluation metrics used (scores greater than 4 in all metrics), that is, they are cohesive texts with regard to structure, spelling and clinical coherence.
Authenticity of Format and Structure: the models were assessed on their adherence to the format and structure typical of clinical documents. Sabia-7B and Mistral-7B-v0.2 scored 3.84 and 3.85, respectively, indicating moderate adherence. LlaMA-2-7B demonstrated better consistency with a score of 4.45. Our models, Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-2-7B, achieved the highest scores of 4.60 and 4.62, respectively. These models also had a significant number of perfect scores (5), reflecting their superior performance in maintaining document authenticity.
Spelling Accuracy: despite Sabia-7B not achieving above 4 score average in Criteria 1 and 3, it demonstrated excellent performance in orthographic correctness, likely due to its training on a large corpus of pt-BR data. In contrast, the Mistral-7B-v0.2 model showed significant difficulties, often generating random texts or words in English. LlaMA-2-7B, although not trained with a large Portuguese corpus, managed to perform well in this criterion. As indicated by the graph, our adjusted LlaMA model (i.e., Clinical-BR-LlaMA-2-7B) further improved upon the already good performance of LlaMA-2-7B. The Mistral-7B-v0.2 model showed a remarkable improvement, with its base model scoring an average of 3.82 and achieving an increase to 4.69, including 30 additional perfect scores (5) in the frequency distribution, indicating excellent performance.
Clinical Coherence: in terms of Clinical Coherence, Sabia-7B scored 3.54 and Mistral-7B-v0.2 scored 3.52, both struggling to maintain logical correlation within the clinical context. LlaMA-2-7B scored 4.19, indicating a better, though not optimal, performance. The models Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-2-7B again led with scores of 4.45 and 4.46, respectively, demonstrating their ability to generate clinically coherent content. Frequency analysis revealed that these models had the highest number of perfect scores (5), underscoring their superior coherence.
4.2 Error Analysis
In this section, we perform an error analysis in the task of generating clinical texts, both from the models we trained and the original baseline models. Table 4 presents some examples of generated clinical texts that obtained low scores for one or more evaluation criteria.
All the evaluated models have demonstrated a behavior in which they repeat certain words or sequences multiple times in succession, like on error #2. This issue can probably be solved by adjusting some inference parameters. For example, when applying the Temperature = 0.1 and Repetition Penalty = 1.2, the output for error #2 would be “por sensação de pressão no peito. A doença foi diagnosticada há mais de um ano atrás, mas o paciente não tomou nenhuma medicação para tratamento. O exame clínico revelou: pressão arterial sistólica de 140 mmHg; frequência cardíaca regular de 76 bpm; pulso normal; auscultação cardiovascular sem alterações\(\ldots \)”.
Although clinical texts are known for not having a standard formal structure, as each physician or institution may establish a different format, we expect a clinical note to contain at least a partial description of the patient’s condition or visit, either in narrative or semi-structured form. Therefore, some notes generated contained an output more similar to a test question or teaching documentation than a clinical narrative (as shown in error #1). This was a more prevalent behavior of the Sabia-7B model, most likely due to its training data related to teaching exams. Regarding the problems in the spelling metric, most of the issues came from generating an output in English instead of Portuguese (like on error #3), being a more recurrent problem in the Mistral-7B-v0.2 model.
As for the clinical coherence dimension, the most complex for models to achieve maximum scores, most of the problems are associated with the use of medical terms that would make sense to be in that text section, but not for the patient’s reported condition. For example, error #4 shows a text that suggests the use of atenolol, a drug widely used to treat hypertension, for the treatment of dyslipidemia. This is an expected problem, given that the standard behavior of models based on vector representations interprets all terms related to drugs as similar, since they are always used in similar contexts. We also had clinical coherence problems, in which the generated output uses keywords related to the patient’s context, but the construction of sentences with these words does not make sense, as in error #5, where the word “dialysis” makes sense, since the patient has chronic kidney disease, but the presented sentence does not make sense from a clinical point of view.
5 Discussion and Future Work
In this study, we developed and evaluated two medical large language models, Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2, tailored for pt-BR. Our findings showed significant improvements in generating synthetic clinical text, particularly in structure, orthographic accuracy, and clinical coherence.
These improvements are due to targeted pre-training and fine-tuning on clinical narratives using Low-Rank Adaptation (LoRA) for efficient adaptation. The Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2 models produced outputs that adhered to clinical document norms, maintained high spelling accuracy, and generated clinically coherent content.
Clinical-BR-Mistral-7B-v0.2 handled long contexts better due to its larger context window, a critical attribute for long clinical documents. It is worth noting that the Mistral-7B-v0.2 model performed worse than LlaMA-2-7B in all evaluation criteria, maybe due to the low volume of pt-BR data in its training (this information was not made available by the team that developed the model). However, after our training on clinical data, the Clinical-BR-Mistral-7B-v0.2 model performed better or similarly to the Clinical-BR-LlaMA-2-7B model. This may indicate that architectural details of the Mistral-based models better handle adjustments to new contexts in a low volume of data scenario and with the use of PEFT techniques.
Our work focuses on medical LLMs for pt-BR, addressing unique challenges compared to models trained in other languages. This distinction is critical as pt-BR linguistic and clinical terminologies differ significantly, affecting model generalization.
BioMistral and MediTron models, trained on extensive medical corpora like PubMed Central, show performance gains in English medical QA tasks. However, our pt-BR models face fewer resources and less standardized evaluations, complicating direct performance comparisons. GatorTronGPT and OpenBioLLM-70B handle extensive clinical data in English, showcasing improvements in biomedical NLP tasks. In contrast, our approach optimizes limited resources while maintaining high performance in pt-BR clinical narratives, emphasizing computational efficiency with techniques like LoRA. This aspect is crucial for making advanced LLMs accessible to research communities with limited computational infrastructure.
Evaluation protocols often involve well-defined benchmarks in English. BioMistral’s evaluation includes multilingual medical QA tasks, reflecting its capabilities. Our evaluation metrics are tailored to pt-BR, ensuring relevance but limiting direct comparisons with English-centric models.
As another limitation in the evaluation protocol of our project, we can mention that the use of medical students may not have been an optimal solution, especially when evaluating the clinical cohesion of the notes, as students have limited expertise regarding some conditions and treatments. Moreover, the length of the clinical notes, a critical factor in generating synthetic health data, was not included in our structure or cohesion assessment. Our primary aim was to identify the impact of pt-BR clinical training data on model performance, even in small volumes, compared to existing models. Consequently, our models are not recommended for generating fully coherent synthetic data. Despite high scores, we did not consider the note length or completeness, leading models to sometimes generate only a few words post-input. Additionally, using other clinical notes as input references may complicate synthetic data generation in environments with limited real clinical data.
While our models show promising results in pt-BR, future work should aim to fine-tune and evaluate the models in clinical downstream tasks, and align evaluation protocols more closely with international benchmarks to facilitate better comparisons. In addition to also building more specific benchmarks for the context of clinical texts, which have a very peculiar format and structure, differing greatly from scientific articles or medical licensing exams. Furthermore, expanding our dataset and incorporating more diverse clinical narratives can enhance model robustness and generalization.
Training a model with medical records data tends to perform better in tasks related to data extraction, interpretation, and generation in this context. However, we understand that several tasks in the medical field involve medical reasoning, and much of the knowledge needed for the model to perform this type of inference is found in biomedical texts and specific question-answering datasets. In this context, we intend to expand our models to the biomedical context as well, so that they are able to deal well with all types of medical data.
By addressing these differences, we highlight the unique contributions of our work and set the stage for future advancements in developing and evaluating medical LLMs across diverse languages and resource settings.
6 Conclusion
This study focused on developing and evaluating two medical large language models, Clinical-BR-LlaMA-2-7B and Clinical-BR-Mistral-7B-v0.2, made for medical Brazilian Portuguese context. These models demonstrated significant improvements over baseline models in generating synthetic clinical text, particularly in terms of Authenticity of Format and Structure, Spelling Accuracy, and Clinical Coherence, largely due to the efficient use of the LoRA technique and continued pretraining on a dataset of clinical notes.
The models’ performance underscores the potential for integrating resource-efficient medical LLMs into clinical practice, facilitating the generation of clinical notes and supporting medical research in Portuguese-speaking contexts. However, limitations such as the subjectivity of the evaluation protocol, the expertise of the evaluators, and the scope of the dataset highlight areas for further enhancement.
Future research should aim to perform the models fine-tuning for specific tasks, align evaluation protocols with international benchmarks, build specific benchmarks for clinical texts, and expand datasets to include diverse clinical narratives and biomedical texts. These efforts will help improve model robustness and generalization, enabling the models to perform better in various medical tasks.
Our work highlights the feasibility of creating resource-efficient medical LLMs for pt-BR, paving the way for their broader adoption in healthcare. Continued improvements and expansions will contribute to more accurate and efficient healthcare solutions, benefiting both patients and medical professionals.
References
Almeida, T.S., Abonizio, H., Nogueira, R., Pires, R.: Sabiá-2: a new generation of Portuguese large language models (2024)
Ankit Pal, M.S.: Openbiollms: advancing open-source large language models for healthcare and life sciences (2024). https://huggingface.co/aaditya/OpenBioLLM-Llama3-70B
Brown, T.B., Mann, B., et al.: Language models are few-shot learners (2020)
Chen, Z., Cano, A.H., et al.: Meditron-70b: scaling medical pretraining for large language models (2023)
Clusmann, J., et al.: The future landscape of large language models in medicine. Commun. Med. 3(1) (2023)
Conneau, A., et al.: Unsupervised cross-lingual representation learning at scale (2020)
Devlin, J., Chang, M.W., Lee, K., Toutanova, K.: BERT: pre-training of deep bidirectional transformers for language understanding (2019)
Dias, H., Ulbrich, A.H.D.P.d.: Brateca (Brazilian tertiary care dataset): a clinical information dataset for the Portuguese language (2022). https://physionet.org/content/brateca/1.1/
Garcia, G.L., et al.: Introducing bode: a fine-tuned large language model for Portuguese prompt-based task (2024)
Goldberger, A.L., et al.: Physiobank, physiotoolkit, and physionet: components of a new research resource for complex physiologic signals. Circulation 101(23), e215–e220 (2000). [Online]
Hu, E.J., et al.: LoRA: low-rank adaptation of large language models (2021)
Huang, K., Altosaar, J., Ranganath, R.: ClinicalBERT: modeling clinical notes and predicting hospital readmission (2020)
Jiang, A.Q., et al.: Mistral 7b (2023)
Jin, D., Pan, E., Oufattole, N., Weng, W.H., Fang, H., Szolovits, P.: What disease does this patient have? A large-scale open domain question answering dataset from medical exams. Appl. Sci. 11(14), 6421 (2021)
Labrak, Y., Bazoge, A., Morin, E., Gourraud, P.A., Rouvier, M., Dufour, R.: Biomistral: a collection of open-source pretrained large language models for medical domains (2024)
Lee, J., et al.: BioBERT: a pre-trained biomedical language representation model for biomedical text mining. Bioinformatics 36(4), 1234–1240 (2019)
Lopes, F., Teixeira, C., Gonçalo Oliveira, H.: Contributions to clinical named entity recognition in Portuguese. In: Proceedings of the 18th BioNLP Workshop and Shared Task, pp. 223–233. Association for Computational Linguistics, Florence, Italy (2019)
Oliveira, L.E.S.e., et al.: SemClinBr - a multi-institutional and multi-specialty semantically annotated corpus for Portuguese clinical NLP tasks. J. Biomed. Semant. 13(1) (2022)
Peng, C., et al.: A study of generative large language model for medical research and healthcare. NPJ Digit. Med. 6(1) (2023)
Pires, R., Abonizio, H., Almeida, T.S., Nogueira, R.: Sabiá: Portuguese Large Language Models, pp. 226–240. Springer, Cham (2023)
Rasmy, L., Xiang, Y., Xie, Z., Tao, C., Zhi, D.: Med-BERT: pre-trained contextualized embeddings on large-scale structured electronic health records for disease prediction (2020)
Schneider, E.T.R., et al.: CardioBERTpt: transformer-based models for cardiology language representation in Portuguese. In: 2023 IEEE 36th International Symposium on Computer-Based Medical Systems (CBMS), pp. 378–381 (2023)
Schneider, E.T.R., et al.: BioBERTpt - a Portuguese neural language model for clinical named entity recognition. In: Proceedings of the 3rd Clinical Natural Language Processing Workshop, pp. 65–72. Association for Computational Linguistics, Online (2020)
Schneider, E.T.R., de Souza, J.V.A., Gumiel, Y.B., Moro, C., Paraiso, E.C.: A GPT-2 language model for biomedical texts in Portuguese. In: 2021 IEEE 34th International Symposium on Computer-Based Medical Systems (CBMS), pp. 474–479 (2021)
Touvron, H., Martin, L., et al.: Llama 2: open foundation and fine-tuned chat models (2023)
Vaswani, A., et al.: Attention is all you need (2023)
Wu, S., Dredze, M.: Beto, bentz, becas: the surprising cross-lingual effectiveness of BERT (2019)
Yu, H., et al.: Large language models in biomedical and health informatics: a bibliometric review (2024)
Zheng, Y., Gan, W., Chen, Z., Qi, Z., Liang, Q., Yu, P.S.: Large language models for medicine: a survey (2024)
Zheng, Y., Zhang, R., Zhang, J., Ye, Y., Luo, Z., Ma, Y.: Llamafactory: unified efficient fine-tuning of 100+ language models (2024)
Acknowledgments
We would like to thank Comsentimento and the Google for Startups Cloud Program for providing infrastructure and support to develop this project. We acknowledge the use of Generative AI tools, particularly ChatGPT, in the preparation of this article. ChatGPT was employed for tasks such as text revision, paragraph summarization, and enhancing the overall clarity and coherence of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Ethics declarations
Disclosure of Interests
The authors have no competing interests to declare that are relevant to the content of this article.
Rights and permissions
Copyright information
© 2025 The Author(s), under exclusive license to Springer Nature Switzerland AG
About this paper
Cite this paper
de Souza Pinto, J.G., Rodrigues de Freitas, A., Martins, A.C.G., Sawazaki, C.M.R., Vidal, C., Silva e Oliveira, L.E. (2025). Developing Resource-Efficient Clinical LLMs for Brazilian Portuguese. In: Paes, A., Verri, F.A.N. (eds) Intelligent Systems. BRACIS 2024. Lecture Notes in Computer Science(), vol 15415. Springer, Cham. https://doi.org/10.1007/978-3-031-79038-6_4
Download citation
DOI: https://doi.org/10.1007/978-3-031-79038-6_4
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-031-79037-9
Online ISBN: 978-3-031-79038-6
eBook Packages: Computer ScienceComputer Science (R0)